The impact of zinc oxide nanoparticles on phosphorus removal and the microbial community in activated sludge in an SBR†
S. T. Wang*,
W. Q. Wang,
Z. R. Zhang and
H. You
State Key Laboratory of Urban Water Resource and Environment, Harbin Institute of Technology, Harbin 150090, China. E-mail: wshutao@hit.edu.cn
First published on 5th October 2016
Abstract
The impact of zinc oxide nanoparticles (ZnO NPs) on phosphorus removal and the microbial community was investigated after long-term exposure to a simulated SBR. ZnO NPs at low concentrations (1, 5 mg L−1) did not exert an obvious impact on phosphorus removal, but the removal efficiency decreased by 20% and 75%, respectively, in the presence of 10 and 20 mg L−1 ZnO NPs. At these concentrations, ZnO NPs inhibited the activity of exopolyphosphatase (PPX) and polyphosphate kinase (PPK), and extracellular polymeric substance (EPS) production and destroyed the integrity of cell membranes. High-throughput sequencing analysis revealed that 20 mg L−1 ZnO NPs led to an obvious decrease in the community diversity and a shift in the community structure. In addition, the exposure to 10 and 20 mg L−1 ZnO NPs resulted in a decrease in the proportion of Betaproteobacteria and Rhodocyclales (polyphosphate accumulation organisms, PAOs) in the community and led to a promotion in the growth of Alphaproteobacteria and Gammaproteobacteria (glycogen accumulating organisms, GAOs). These findings meaningfully show the adverse effects of NPs on activated sludge in wastewater treatment.
1. Introduction
Large quantities of nanomaterials, zinc oxide (ZnO) for example, have been used in many commercial, industrial and consumer products. Inevitably, some of the ZnO nanoparticles (ZnO NPs) enter industrial and domestic wastewater treatment plants (WWTPs). NPs can be more toxic than larger particles of the same composition because of their large specific surface area and unique size effect.1–3 Many studies have confirmed that ZnO NPs are present in sewage sludge and effluents. According to a report issued by the USEPA in 2009, an examination of 84 WWTPs showed that the zinc content in WWTP biosolids was 8.55 g per kg-SS.4 Investigations in China in 2011 (139 WWTPs in total) and in 2009 (107 WWTPs in total) showed that the average concentration of Zn in biosolids was 1.03 g per kg-SS, and the maximum concentration was 9.14 g per kg-SS.5,6Phosphorus removal is normally achieved through enhanced biological phosphorus removal (EBPR). Under anaerobic conditions, polyphosphate accumulation organisms (PAOs) take up wastewater organic compounds (such as short chain fatty acids) and store them as intracellular polyhydroxyalkanoates (PHA) by utilizing energy and reduction equivalents (NADH2). Polyphosphate hydrolysis, which results in an anaerobic phosphorus release, and glycogen degradation generate energy and NADH2, respectively. Then, the stored PHA is oxidized by PAOs to produce energy for cell growth, maintenance and phosphorus uptake in the subsequent aerobic stage. As phosphorus aerobic uptake is greater than the anaerobic release, a net phosphorus removal is achieved by wasting excess sludge.
It is well known that NPs have antibacterial abilities.7 Concern about whether ZnO NPs will negatively impact the function of microorganisms in WWTPs is therefore raised. Studies have addressed this issue8–10 reported that the increase of Ag NPs showed no significant influence on EBPR in both short-term and long-term experiments. Phosphorus removal, however, was decreased by a sudden increase in Ag+ concentration, but it was gradually recovered after long-term culture.
It was also found that phosphorus removal efficiency was decreased to 48.8% at 1 mg L−1 of Ag+, and no net phosphorus was removed at >2 mg L−1 of Ag+ in batch tests.11 The uptake of wastewater carbon sources and the anaerobic and aerobic transformations of phosphorus, PHA and glycogen were inhibited by Ag+ rather than Ag NPs. Ag+ showed stronger toxicity to PAOs than to glycogen accumulating organisms (GAOs).
Zheng et al.12 found that phosphorus removal was insensitive to 1 and 50 mg L−1 SiO2 NPs after either acute or chronic exposure. The critical factors related to biological phosphorus removal were not significantly changed, such as the activities of exopolyphosphatase (PPX) and polyphosphate kinase (PPK) and the intracellular transformations of PHA and glycogen. It was found that 1 and 50 mg L−1 TiO2 NPs had no acute effects on wastewater nitrogen and phosphorus removal after short-term exposure (1 d). Concentrations of 50 mg L−1 TiO2 NPs did not affect biological phosphorus removal after long-term exposure (70 d), even though total nitrogen (TN) removal efficiency was decreased from 80.3% to 24.4%.12
ZnO NPs exhibited significant toxicity to the activated sludge at low concentrations compared to other NPs13 (Mu et al., 2011). Unlike many other NPs, the slight solubility of ZnO NPs may have contributed to its higher toxicity.14 It was found that ZnO NPs caused poor settleability of the activated sludge and a significant decrease in the removal of nitrogen and phosphorus over time. The bacterial community in the activated sludge also became less diverse after exposure to ZnO NPs. It was revealed that although the COD and NH4+–N removal efficiencies were not reduced by the addition of ZnO NP or only slightly reduced, the functional bacterial community changed remarkably. Zheng et al.12 conducted short-term exposure study to determine whether ZnO NPs caused adverse impacts on biological nitrogen and phosphorus removal in the unacclimated anaerobic-low dissolved oxygen SBR. Their findings showed that, compared to the absence of ZnO NPs, the presence of 10 and 50 mg L−1 ZnO NPs decreased the TN removal efficiencies from 81.5% to 75.6% and 70.8%, respectively, and the effluent phosphorus concentrations increased from undetectable to 10.3 and 16.5 mg L−1, respectively, which were higher than the influent phosphorus (9.8 mg L−1). It was found that the inhibition of nitrogen and phosphorus removal induced by higher concentrations of ZnO NPs was mainly due to the release of zinc ions from ZnO NPs dissolution and the increase of reactive oxygen species (ROS) production, which caused decreased activities of nitrate reductase, PPX, and PPK. Chen et al.10 focused on the effects of ZnO NPs on the functional bacterial community in wastewater treatment. They found that although the NH4+–N removal efficiency was not or was only slightly reduced, the denitrification-related species were inhibited by 5 mg L−1 ZnO NPs, including Diaphorobacter species, Thauera species and those in the Sphaerotilus-Leptothrix group. However, the bacteria related to sludge bulking (Haliscomenobacter hydrossis), heavy metal resistance (Zoogloea ramigera) and biosorption (Methyloversatilis universalis) were increased by ZnO NPs treatment. It was also concluded that ZnO NPs slightly decreased TN removal. Moreover, in the simulated SBR processes, continuous input of ZnO NPs into the wastewater reduced NH4+–N removal by inhibiting the respiration of nitrifying microorganisms.15
A few studies reported the potential effects of ZnO NPs on biological phosphorus removal and microbial community. The focus of this study was to determine the effects of ZnO NPs on biological phosphorus removal and the microbiological community after long-term exposure by using high-throughput sequencing technique in the anaerobic–aerobic sequencing batch reactor (SBR). The activities of some key enzymes related to biological phosphorus removal and EPS production and plasma membrane integrity were further measured to explore the potential effects of ZnO NPs on activated sludge.
2. Material and methods
2.1. Nanoparticle suspension
A commercially produced ZnO NP suspension (1.7 g L−1, <100 nm particle size) was purchased from Sigma-Aldrich (St Louis, MO, USA). Before use, the ZnO NP suspension was prepared by dispensing 1 mL of ZnO NPs (1.7 g L−1) into 1 L of deionized water, followed by 1 h of sonication (25 °C, 250 W, 40 kHz). The average particle size in the stock suspension was measured to be in the range from 30 to 60 nm by dynamic light scattering (DLS) analysis via a Malvern Autosizer (4700, Malvern Instruments, UK). According to transmission electron microscopy (TEM) and scanning electron microscopy (SEM), the ZnO NPs were spherical.2.2. SBR operation
The reactor was inoculated with conventional activated sludge taken from the Harbin Taiping Municipal Wastewater Treatment Plant, Harbin, China. All of the reactors were fed with synthetic wastewater containing the following components: NH4+–N (NH4Cl), 30–50 mg L−1; COD (sucrose), 350–450 mg L−1; KH2PO4, 30 mg L−1; MgSO4·7H2O, 50 mg L−1 and CaCl2·2H2O, 10 mg L−1. The activated sludge was successfully cultured with the synthetic wastewater.The activated sludge was cultured in the SBR with a working volume of 4 L, which was operated to achieve biological nitrogen removal. The SBR was operated at 25–28 °C with three cycles each day. Each cycle (8 h) consisted of 5.6 h of aeration, followed by 1 h of settling, 15 min for decanting and 1 h idle. The influent pH was adjusted to 7.5 by adding NaOH, NaHCO3 and HCl. Air was provided intermittently by using an on/off controller with an online DO detector to maintain DO at an appropriate level. Sludge was wasted to keep the solids retention time (SRT) at approximately 22 d to maintain the ratio of mixed liquor volatile suspended solid (MLVSS) to mixed liquor suspended solids (MLSS). Namely MLVSS/MLSS, at 0.75. The reactor was constantly mixed with a stirrer except during the settling, decanting, and idle periods. After approximately three months, stable removal efficiency of phosphorus (>90.0%) was achieved.
2.3. ZnO NPs exposure to activated sludge
One control and four test concentrations (1, 5, 10 and 20 mg L−1) of ZnO NPs were examined in the exposure experiments. The environmentally relevant concentrations of total NPs in WWTPs are from 1 to 5 mg L−1 accordingly.16 Because the environmental release of NPs might increase with large-scale manufacturing, the potential effects of higher concentrations of 10 and 20 mg L−1 ZnO NPs were also investigated. Two SBR reactors with a working volume of 4 L each were used to conduct the experiments. Each cycle (8 h) consisted of 2 h anaerobic and 4 h aerobic periods, followed by 1 h of settling, 10 min for decanting and 50 min idle. Sludge was wasted to keep the solids retention time (SRT) at approximately 22 d. Each SBR reactor was set up in triplicate to ensure reliable results. After the addition of ZnO NPs, the trend of the changes in phosphorus during each operating cycle (8 h) and long-term period (10 d) were all measured.2.4. Analytical methods
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 10 min at 6–10 °C, and the precipitate was resuspended to 30 mL. This process was repeated twice. The pretreated activated sludge was dissolved in 15 mL of TE buffer solution. Bulk genomic DNA was extracted using sodium dodecyl sulfate (SDS) and hexadecyltrimethyl ammonium bromide (CTAB), and the products were examined by agarose (1% w/v) gel electrophoresis in Tris/borate/EDTA buffer (TBE).
000g for 10 min at 6–10 °C, and the precipitate was resuspended to 30 mL. This process was repeated twice. The pretreated activated sludge was dissolved in 15 mL of TE buffer solution. Bulk genomic DNA was extracted using sodium dodecyl sulfate (SDS) and hexadecyltrimethyl ammonium bromide (CTAB), and the products were examined by agarose (1% w/v) gel electrophoresis in Tris/borate/EDTA buffer (TBE).Total genomic DNA from microbial samples was first extracted and purified using a PowerSoil DNA Isolation Kit (MoBio Laboratories, Inc., Carlsbad, CA) according to manufacturer protocol and then detected with 1% agarose gel electrophoresis and stored at −20 °C until use. 16 S rRNA genes were amplified with barcoded primers 515F (5′-GTGCCAGCMGCCGCGG-3′)/907R (5′-CCGTCAATTCMTTTRAGTTT-3′) as follows: initial denaturation at 95 °C for 3 min, 28 cycles of 95 °C for 30 s, 55 °C for 30 s, 72 °C for 45 s, followed by a final extension 72 °C for 10 min and 10 °C until halted by user. Each 20 μL reaction mixture consisted of 4 μL of 5× FastPfu buffer, 2 μL of deoxynucleoside triphosphate (dNTP) mix (2.5 mM each), 10 ng of template DNA, 0.8 μL of 5 μM barcode primer 515F, 0.8 μL of 5 μM barcode primer 907R, with double-distilled H2O supplemented to 20 μL. PCR products were visualized on an agarose gel and mixed proportional with each other according to mass prior to sequencing on an Illumina Miseq benchtop sequencer using pair-end 250 bp kits at Shanghai (majorbio), China.
The raw sequences were optimized, and low quality sequences were removed using the Mothur (http://www.mothur.org). Briefly, Mothur was used to trim barcode and primer sequences and to eliminate sequences shorter than 200 bp, with one or more ambiguous bases and with a quality score inferior to 25. Sequences were clustered into Operational Taxonomic Units (OTUs) at 97% sequence similarity by using Mothur. Species richness, diversity indices (i.e., observed OTUs, Chao1 estimator, Shannon index, Simpson index and abundance-based coverage estimator (ACE)), and rarefaction curves were obtained using Mothur, at a 3% dissimilarity cutoff. To compare the community diversity between samples based on phylogenetic information, the Fast UniFrac online tool (http://unifrac.colorado.edu/) was used to estimate the weighted UniFrac metric and to carry out principal coordinate analysis (PcoA). Moreover, a heatmap was implemented through the use of R packages heatmap (http://www.r-project.org/). The 16S rRNA gene sequences were deposited in the NCBI Sequence Read Archive under accession number.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g for 5 min. Then, the supernatant was seeded on a 96-well plate, followed by the addition of 50 μL of substrate mix (Tiangen). After incubation at room temperature for 30 min in the dark, 50 μL of stop solution (Tiangen) was added to each well, and the absorbance was recorded at 490 nm using a microplate reader (BioTek).
000 g for 5 min. Then, the supernatant was seeded on a 96-well plate, followed by the addition of 50 μL of substrate mix (Tiangen). After incubation at room temperature for 30 min in the dark, 50 μL of stop solution (Tiangen) was added to each well, and the absorbance was recorded at 490 nm using a microplate reader (BioTek).2.5. Statistical analysis
All of the tests were performed in triplicate, and the results were expressed as the mean ± standard deviation. Analysis of variance (ANOVA) was used to examine the significance of the results, and p < 0.05 was considered to be statistically significant.3. Results and discussion
3.1. Effects of ZnO NPs on biological phosphorus removal
The effect of ZnO NPs on the content of PHA and glycogen during biological phosphorus removal in SBR is shown in Fig. 1A and B. It indicates that the content (%/MLVSS) of PHA and glycogen were both similar to those of the control when the exposure concentration of ZnO NPs was 1 mg L−1. Compared to the control, when the concentrations of ZnO NPs were increased to 5, 10 and 20 mg L−1, the content of PHA was increased from 7.01% to 7.33%, 7.42% and 7.59%, respectively, at the end of the anaerobic stage, and it was increased from 2.90% to 3.07%, 3.22% and 3.39% at the end of aerobic stage, respectively. It indicates that higher concentrations of ZnO NPs could increase PHA content. As shown in Fig. 1B, compared to the control, the content of glycogen decreased from 9.51% to 9.11%, 8.58% and 8.14%, respectively, at the end of anaerobic stage at the ZnO NPs concentrations of 5, 10 and 20 mg L−1. However, glycogen content increased from 12.02% to 12.44%, 12.95% and 13.37% at the end of aerobic stage, respectively. It indicates that ZnO NPs at higher concentrations could improve the activity of PAOs by promoting the decomposition and synthesis of glycogen.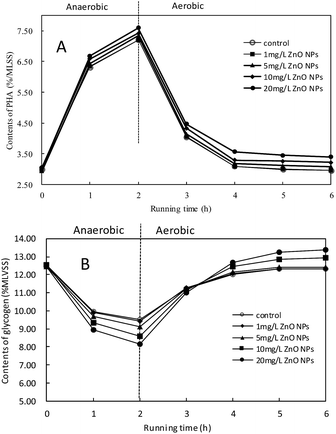 | ||
| Fig. 1 Effects of ZnO NPs on the contents of PHA and glycogen in activated sludge in SBR during one cycle. PHA (A); glycogen (B). | ||
It is reported that the increase in PHA contents was due to synthesis of PHV,18 which was generated during the decomposition of glycogen by GAOs. The increase in PHV proved the increase in the activity of GAOs, which is in accordance with the changes in glycogen noted above. Accordingly, PAOs lost the competitive advantage against GAOs, leading to the adverse effect of higher concentrations of ZnO NPs on phosphorus removal.
The effect of ZnO NPs on phosphorus release in the anaerobic stage and phosphorus removal in the effluent was investigated as shown in Fig. 2A and B. It can be seen that in the presence of 1 and 5 mg L−1 ZnO NPs, the release of SOP in the anaerobic stage was relatively stable and similar to that of the control.
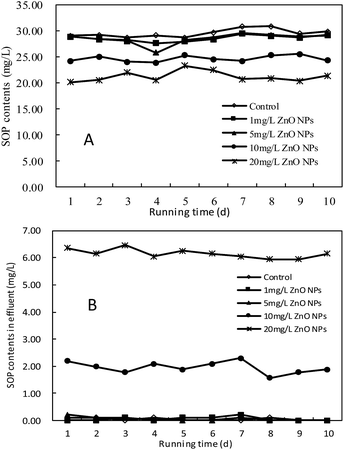 | ||
| Fig. 2 Effect of ZnO NPs on phosphorus release in the anaerobic stage and phosphorus removal of the SBR. | ||
The phosphorus removal efficiencies were almost 100%, which indicated that low concentration of ZnO NPs (1, 5 mg L−1) had no significant adverse effect on phosphorus removal. However, the releases of SOP in the anaerobic stage were decreased by 20% and 27%, respectively when 10 and 20 mg L−1 ZnO NPs were exposed to SBR. Additionally, the effluent SOP increased from undetectable to 1.95 ± 0.3 mg L−1 and 6.20 ± 0.2 mg L−1, respectively. The above results show that 10 and 20 mg L−1 ZnO NPs led to the poor performance of biological phosphorus removal.
3.2. Effects of ZnO NPs on activity of the activated sludge
Phosphorus removal is directly related to the activity of PPX and PPK. As shown in Fig. 3A, with the increase of ZnO NPs concentrations, the inhibition of the activities of PPX and PPK was strengthened. For 1, 5, 10 and 20 mg L−1 of ZnO NPs, the ratios of inhibition to PPX were 1.21%, 3.25%, 10.27% and 28.05%, respectively, and the ratios of inhibition to PPK were 1.32%, 4.13%, 11.15% and 30.13%, respectively. The numerical difference of the enzyme activity was very limited because the measured unit of enzyme activity is very small (0.006–0.009 OD per mg proteins). Therefore, it suggests that the addition of 1 and 5 mg L−1 ZnO NPs did not cause inhibitory effects on PPX and PPK. Whereas, 10 and 20 mg L−1 ZnO NPs inhibited the activities of PPX and PPK significantly by affecting the normal function of PAOs.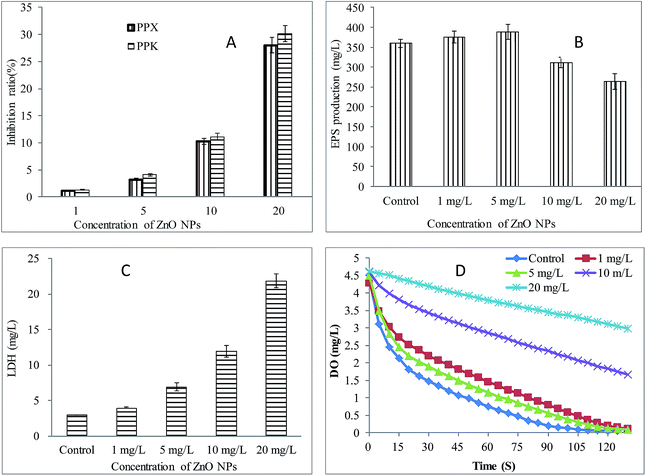 | ||
| Fig. 3 Effects of ZnO NPs on the activity of the activated sludge. (A) Activity of PPX and PPK; (B) EPS production; (C) LDH level; (D) respiration rate. | ||
Extracellular polymeric substances (EPS) play an important role in protecting cells. The results indicate that 1 and 5 mg L−1 ZnO NPs had no adverse effect on EPS yield in activated sludge system, and in fact, EPS yield was increased by 4.4% and 7.8%, respectively (Fig. 3B). The addition of 10 and 20 mg L−1 ZnO NPs reduced the production of EPS, and the reduction rate was 13.3% and 27.7%, respectively. EPS production and cell metabolism are closely related. The addition of higher concentrations of ZnO NPs (10, 20 mg L−1) reduced the yield of EPS, suggesting ZnO NPs resulted in inhibition to bacterial metabolism and degradation of external matrix, and the higher the concentration, the greater inhibition.
The effect of ZnO NPs on cell membrane integrity of activated sludge was studied by determining LDH release. As shown in Fig. 3C, the exposure to 1 and 5 mg L−1 ZnO NPs caused a slight increase in LDH level, indicating that low concentrations of ZnO NPs caused a little damage to the cell membrane and little leakage of the cytoplasm. Compared to the control (2.99 U L−1), however, the exposure to 10 and 20 mg L−1 ZnO NPs led to an increase in LDH level by approximately 300% and 600%, respectively, indicating that higher concentrations of ZnO NPs caused significant damage to the integrity of cell membrane and led to significant leakage of cytoplasm.
The respiration rate of activated sludge was determined by the least squares fitting line. As shown in Fig. 3D, respiratory rate of each system was 77.54, 80.14, 85.53, 66.31 and 46.87 mg O2 per L per h, respectively. The results indicated that the exposure to 1 and 5 mg L−1 ZnO NPs did not cause inhibition to the respiratory rate of activated sludge. On the contrary, the respiration was slightly enhanced. In the presence of 10 and 20 mg L−1 ZnO NPs, the respiration was inhibited by 14.5% and 39.6%, respectively, compared to that of the control. It shows that low concentrations of ZnO NPs (1, 5 mg L−1) could slightly improve the oxygen uptake rate, suggesting that microorganisms could adjust themselves to adapt and resist the toxic impact of ZnO NPs, but 10 and 20 mg L−1 ZnO NPs could obviously inhibit the biological respiration of activated sludge by affecting the normal microbial metabolic activity.
3.3. Effects of ZnO NPs on biological community
These sequences were clustered according to the similarity (0.97), and each group was an operational taxonomic unit (OTU). The VENN diagram (Fig. 4) shows the number of OTUs in different samples. The OTU number of P0, P1, P5, P10 and P20 systems, was 6255, 5029, 5359, 5979 and 4910, respectively.
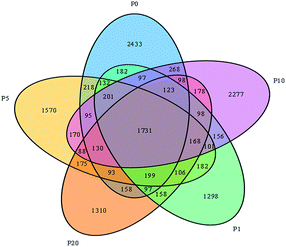 | ||
| Fig. 4 OTU-VEEN diagram based on high-throughput sequencing analysis. P0, P1, P5, P10 and P20 represents 0, 1, 5, 10 and 20 mg L−1 ZnO NPs exposure system, respectively. | ||
The alpha diversity index reflects the microbial community diversity of the activated sludge. Table 1 shows the alpha diversity statistics for the five samples. Chao1 and Ace indices were used to estimate the total biological species in the community by utilizing different calculation methods. The OTU number and the species richness in the samples were positively correlated. Shannon and Simpson indices were used to estimate the microbial diversity in the activated sludge systems. The higher the Shannon index, the higher the diversity of microbial community. In contrast, a lower Simpson index represents a higher diversity of the community. In addition, based on high-throughput sequencing analysis, the richness rarefaction was plotted to indicate the species abundance in the activated sludge (Fig. S2, ESI†).
| Sample | Sequence Number | OTU Number | Shannon | Ace | Chao1 | Coverage | Simpson |
|---|---|---|---|---|---|---|---|
| a Note: P0, P1, P5, P10 and P20 represents 0, 1, 5, 10 and 20 mg L−1 ZnO NPs exposure system, respectively. | |||||||
| P0 | 35507 | 6255 | 7.079310 | 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 520.21 520.21 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 335.34 335.34 |
0.891205 | 0.004368 |
| P1 | 39105 | 5029 | 6.877412 | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 147.58 147.58 |
9797.38 | 0.934152 | 0.004451 |
| P5 | 35258 | 5359 | 6.987293 | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 955.06 955.06 |
11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 919.06 919.06 |
0.915083 | 0.004411 |
| P10 | 30013 | 5979 | 7.142663 | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 135.61 135.61 |
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 321.81 321.81 |
0.874754 | 0.003843 |
| P20 | 35258 | 4910 | 6.890649 | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 037.52 037.52 |
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 044.20 044.20 |
0.926350 | 0.003870 |
As Fig. S2†, 4 and Table 1 show, arranged in descending order, the community diversity of the different systems are P10, P0, P5, P1 and P20 in order. The Simpson index of P0, P1 and P5 were very close; thus, the addition of 1 and 5 mg L−1 ZnO NPs exerted no obvious adverse influence on the community diversity. The addition of 10 mg L−1 ZnO NPs increased the biological community diversity, which indicates that the activated sludge has the ability to self-adjust to resist the toxicity of ZnO NPs. However, the shock of 20 mg L−1 ZnO NPs is too large for microorganism to endure; thus, the biological community diversity is the lowest when exposed to 20 mg L−1 ZnO NPs.
The species were classified and the flora abundances were obtained according to the sequencing results. Based on the abundance matrix, an abundance heatmap was plotted. The heatmap reflected the abundance information of the flora by the shade of the color. As shown in Fig. S3 (ESI†), each row represents one type of flora, each column represents one sample, and the color block represents the abundance of the species. Red indicates a low abundance and blue indicates a high abundance. According to the heatmap, an order level flora abundance was obtained as showed in Fig. S4 (ESI†).
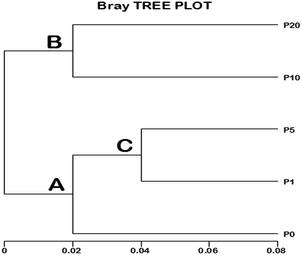
Fig. 5 shows the cluster difference among samples. The left part of the figure represents the clustering tree of the five samples. The more similar the bacteria in these samples, the smaller distance in the cluster tree between samples. From the distance between the five samples in the cluster tree, it could be concluded that the community of the activated sludge system changed after the addition of ZnO NPs. Higher concentrations of ZnO NPs had a stronger effect on the community shift.
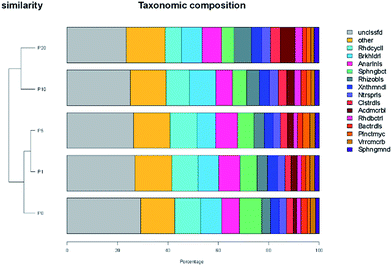 | ||
| Fig. 5 Cluster tree at the order level for the five samples. P0, P1, P5, P10 and P20 represents 0, 1, 5, 10 and 20 mg L−1 ZnO NPs exposure system, respectively. | ||
According to Fig. S3, S4† and Fig. 5, the flora difference among the five samples could be clearly found and the bray tree was plotted as Fig. 6 shows. It can be seen that there were two community branches A and B. branch A contains P0 and a sub-branch C (contains P1 and P5) as well as branch B contains P10 and P20. Theoretically, the closer position represents the more similarity of flora distribution in the bray tree plot. It could be observed that P1 had the similar flora distribution to P5 in sub-branch C and P0 had the similar distribution to the sub-branch C. In addition, in branch B, the flora distribution of P10 was similar to that of P20, but much different to those of P0, P1 and P5. The above analysis proved that the flora distributions were changed when the activated sludge were exposed to ZnO NPs at all the concentrations of 1, 5, 10 and 20 mg L−1 compared to the control. However, the exposure to 1 and 5 mg L−1 ZnO NPs caused slight effect on flora distribution, but the exposure to 10 and 20 mg L−1 ZnO NPs led to significant effect on that.
| Phylum name | Distribution (%) | ||||
|---|---|---|---|---|---|
| P0 | P1 | P5 | P10 | P20 | |
| a Note: P0, P1, P5, P10 and P20 represents 0, 1, 5, 10 and 20 mg L−1 ZnO NPs exposure system, respectively. | |||||
| Proteobacteria | 41.08 | 43.21 | 40.87 | 45.54 | 43.07 |
| Bacteroidetes | 22.13 | 16.91 | 16.23 | 14.38 | 12.37 |
| Chloroflexi | 8.03 | 9.83 | 10 | 7.65 | 9.05 |
| Acidobacteria | 4.98 | 5.82 | 5.39 | 5.35 | 6.21 |
| Firmicutes | 3.79 | 3.73 | 4.39 | 4.89 | 5.71 |
| Nitrospira | 2.93 | 2.89 | 2.81 | 3.53 | 3.39 |
| Verrucomicrobia | 2.84 | 2.87 | 2.96 | 2.03 | 1.87 |
| Actinobacteria | 2.25 | 3.93 | 3.72 | 4.27 | 8.24 |
| Unclassified | 6.06 | 5.2 | 6.01 | 6.05 | 4.84 |
| Class name | Distribution (%) | ||||
|---|---|---|---|---|---|
| P0 | P1 | P5 | P10 | P20 | |
| a Note: P0, P1, P5, P10 and P20 represents 0, 1, 5, 10 and 20 mg L−1 ZnO NPs exposure system, respectively. | |||||
| Betaproteobacteria | 20.2 | 20.36 | 19.58 | 21.32 | 16.39 |
| Alphaproteobacteria | 9.22 | 10.72 | 10.15 | 12.88 | 15.01 |
| Sphingobacteria | 8.85 | 6.77 | 6.52 | 5.64 | 4.8 |
| Gammaproteobacteria | 7.94 | 8.5 | 7.48 | 7.74 | 8.58 |
| Anaerolineae | 7.06 | 8.43 | 8.69 | 6.6 | 7.8 |
| Nitrospira | 2.93 | 2.89 | 2.81 | 3.53 | 3.39 |
| Clostridia | 2.64 | 2.42 | 2.86 | 3.43 | 3.91 |
| Bacteroidia | 2.56 | 1.65 | 2 | 2.17 | 1.58 |
| Eltaproteobacteria | 2.37 | 2.69 | 2.59 | 2.32 | 2.04 |
| Actinobacteria | 2.25 | 3.93 | 3.72 | 4.27 | 8.24 |
| Verrucomicrobiae | 2.13 | 1.95 | 2.14 | 1.47 | 1.28 |
| Acidobacteria_Gp4 | 2.05 | 1.83 | 1.8 | 1.68 | 1.56 |
| Bacteroidetes | 1.34 | 1.01 | 0.98 | 0.78 | 0.67 |
| Planctomycetacia | 1.24 | 1.68 | 2.79 | 1.85 | 1.79 |
| Unclassified | 18.41 | 15.11 | 15.87 | 14.68 | 12.22 |
As Table 2 shows, high-throughput sequencing analysis on the cDNA sequences of the 16S rRNA genes revealed two common phyla, namely, Proteobacteria and Bacteroidetes were retrieved from all five activated sludge systems. Some difference can be observed in the distribution of dominant bacteria in different samples. With the increase of ZnO NPs concentrations, the proportion of Proteobacteria in the sample was generally rising. Proteobacteria was the most important bacteria in the activated sludge system, and many nitrogen fixing bacteria and phosphorus accumulating bacteria are strains of Proteobacteria.19 The addition of ZnO NPs significantly inhibited the activity of Bacteroidetes. It has been reported that Bacteroidetes degrade polycyclic aromatic hydrocarbons (PAHs) and refractory biomacromolecules.20 In the community, the proportion of the bacteria decreased with the increase of the concentrations of ZnO NPs. The proportion of Bacteroidetes was 22.13% in the control test, but it was decreased to 12.37% in the system containing 20 mg L−1 ZnO NPs. There was some difference among the samples in the distribution of different phyla, among which the difference in Actinobacteria were obvious. The activity of Actinobacteria was significantly enhanced by the addition of ZnO NPs, and its proportion was increased on a large scale, from 2.25% to 8.24%. Actinobacteria are heterotrophic bacteria, most strains produce H2S in culture and could deoxidize nitrate into nitrite. The addition of ≤20 mg L−1 of ZnO NPs may promote denitrification, which is consistent with the previous findings.15 Adding the experimental concentrations of ZnO NPs caused minimal impact on the abundance of Nitrospira.
As shown in Tables 3 and 4, in the most abundant phylum Proteobacteria, the beta subclass was particularly predominant, accounting for approximately 20%. In the predominant class Betaproteobacteria, the Rhodocyclales was predominant.
| Order name | Distribution (%) | ||||
|---|---|---|---|---|---|
| P0 | P1 | P5 | P10 | P20 | |
| a Note: P0, P1, P5, P10 and P20 represents 0, 1, 5, 10 and 20 mg L−1 ZnO NPs exposure system, respectively. | |||||
| Rhodocyclales | 10.39 | 10.39 | 10.64 | 9.27 | 6.42 |
| Alphaproteobacteria | 9.22 | 10.72 | 10.15 | 12.88 | 15.01 |
| Sphingobacteriales | 8.85 | 6.77 | 6.52 | 5.64 | 4.8 |
| Burkholderiales | 8.24 | 8.23 | 7.37 | 10.37 | 8.18 |
| Anaerolineales | 7.06 | 8.43 | 8.69 | 6.6 | 7.8 |
| Rhizobiales | 3.49 | 4.09 | 4.14 | 5.15 | 6.9 |
| Xanthomonadales | 3.41 | 4.07 | 3.5 | 3.82 | 4.25 |
| Nitrospirales | 2.93 | 2.89 | 2.81 | 3.53 | 3.39 |
| Clostridia | 2.64 | 2.42 | 2.86 | 3.43 | 3.91 |
| Bacteroidales | 2.56 | 1.65 | 2 | 2.17 | 1.58 |
| Verrucomicrobiales | 2.13 | 1.95 | 2.14 | 1.47 | 1.28 |
| Rhodobacterales | 1.64 | 1.62 | 1.67 | 2.6 | 2.91 |
| Sphingomonadales | 1.42 | 1.82 | 1.65 | 1.78 | 1.99 |
| Acidimicrobiales | 1.4 | 2.48 | 2.37 | 3.04 | 5.8 |
| Planctomycetales | 1.24 | 1.68 | 2.79 | 1.85 | 1.79 |
| Myxococcales | 1.1 | 1.52 | 1.54 | 1.4 | 1.12 |
| Unclassified | 29.18 | 26.86 | 26.39 | 25.1 | 23.52 |
It was concluded that organisms related to the Rhodocyclus-like group within the Proteobacteria beta subclass are the most dominant species responsible for biological phosphorus removal in SBR,21 which is consistent with these experimental results. The proportion of Betaproteobacteria and Rhodocyclales was decreased in systems with the addition of 10 and 20 mg L−1 ZnO NPs, which was consistent with the poor phosphorus removal efficiency. The bacteria abundance results show ZnO NPs played a role in promoting the growth of Alphaproteobacteria and Gammaproteobacteria. Researcher have noted that in a deteriorated biological phosphorus removal reactor, the most dominant sequences were affiliated with the alpha and beta subclass of the Proteobacteria, which were considered as GAOs. It indicates that ZnO NPs improved the growth of GAOs to a certain degree. In biological phosphorus removal system, there is competition between PAOs and GAOs for the carbon substrate. Consistent with these experimental results, activated sludge exposed to 10 and 50 mg L−1 ZnO NPs showed greater variations in total PHA and glycogen in both anaerobic and low DO stages, and the higher transformation of glycogen indicates that the metabolism of GAOs might be activated.
4. Conclusions
Additions of 1 and 5 mg L−1 ZnO NPs showed no obvious effect on the phosphorus removal efficiency after long-term exposure. However, the presence of 10 and 20 mg L−1 ZnO NPs inhibited the average anaerobic SOP release and the phosphorus removal of SBR was decreased. It was revealed that the exposure to 10 and 20 mg L−1 ZnO NPs caused a significant inhibition to the activity of PPX and PPK, and EPS production, respiration rate as well as destroyed the integrity of cell membrane. High-throughput sequencing analysis showed significant difference in the microbial community at the phylum, class and order levels in the presence of 10 and 20 mg L−1 ZnO NPs. A shift occurred in the abundance of the dominant phylum (Proteobacteria, Bacteroidetes and Actinobacteria). The exposure to 10 and 20 mg L−1 ZnO NPs resulted in a decrease in the proportion of Betaproteobacteria and Rhodocyclales (PAOs) in the community and led to a promotion in the growth of Alphaproteobacteria and Gammaproteobacteria (GAOs). This explains the poor phosphorus removal efficiency caused by ZnO NPs.Acknowledgements
The Project was sponsored by the Scientific Research Foundation for the Returned Overseas Chinese Scholars, State Education Ministry, China. This work was also supported by the Key Laboratory of Urban Water Resource and Environment of Harbin institute of technology, China (ES201608).References
- A. Nel, T. Xia and L. Mädler, Science, 2006, 311, 622–627 CrossRef CAS PubMed.
- M. Alaraby, A. Hernández and R. Marcos, Nanotoxicology, 2016, 10, 749–760 CrossRef CAS PubMed.
- S. Jafarirad, M. Mehrabi and B. Divband, Mater. Sci. Eng., C, 2016, 59, 296–302 CrossRef CAS PubMed.
- J. Yang, G. H. Guo and T. B. Chen, China Water Wastewater, 2009, 25, 122–124 CAS.
- X. Mei, Z. Wang and X. Zheng, J. Membr. Sci., 2014, 451, 169–176 CrossRef CAS.
- X. Ma, H. Weng and J. Zhang, China Environ. Sci., 2011, 31, 1306–1313 CAS.
- L. Zhang, Y. Jiang, Y. Ding, M. Povey and D. York, J. Nanopart. Res., 2007, 9, 479–489 CrossRef CAS.
- S. Wang, M. C. Gao, Z. L. She, D. Zheng, C. J. Jin, L. Guo, Y. G. Zhao, Z. W. Li and X. J. Wang, Bioresour. Technol., 2016, 216, 428–436 CrossRef CAS PubMed.
- L. Z. Miao, C. Wang, J. Hou, P. F. Wang, Y. H. Ao, Y. Li, N. Geng, Y. Yao and B. W. Lv, Bioresour. Technol., 2016, 216, 537–544 CrossRef CAS PubMed.
- J. Chen, Y. Q. Tang, Y. Li, Y. Nie, L. L. Hou, X. Q. Li and X. L. Wu, Chemosphere, 2014, 104, 141–148 CrossRef CAS PubMed.
- Y. G. Chen, H. Chen, X. Zheng and H. Mu, J. Hazard. Mater., 2012, 239–240, 88–94 CrossRef CAS PubMed.
- X. Zheng, Y. G. Chen and R. Wu, Environ. Sci. Technol., 2011, 45, 7284–7290 CrossRef CAS PubMed.
- H. Mu, Y. G. Chen and N. Xiao, Bioresour. Technol., 2011, 102, 10305–10311 CrossRef CAS PubMed.
- G. Liu, D. Wang, J. Wang and C. Mendoza, Sci. Total Environ., 2011, 409, 2852–2857 CrossRef CAS PubMed.
- S. T. Wang, S. P. Li, W. Q. Wang and H. You, RSC Adv., 2015, 5, 67335–67342 RSC.
- Y. G. Chen, Y. L. Su, X. Zheng, H. Chen and H. Yang, Water Res., 2012, 46, 4379–4386 CrossRef CAS PubMed.
- Q. X. Wen, Z. Q. Chen, C. Y. Wang and N. Q. Ren, J. Environ. Sci., 2012, 24, 1744–1752 CrossRef CAS.
- Y. Y. Wang, J. J. Geng and Z. J. Ren, J. Chem. Technol. Biotechnol., 2013, 88, 1228–1236 CrossRef CAS.
- O. Henriet, C. Meunier, P. Henry and J. Mahillon, Bioresour. Technol., 2016, 211, 298–306 CrossRef CAS PubMed.
- J. Zhan, F. Y. Ji, X. Xu, X. Y. Xu, Y. K. Chen and Q. Li, Biodegradation, 2014, 25, 777–786 CrossRef PubMed.
- H. Chen, X. Zheng, Y. G. Chen and H. Mu, RSC Adv., 2013, 3, 9835–9842 RSC.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c6ra19486a |
| This journal is © The Royal Society of Chemistry 2016 |