Open Access Article
Open Access ArticleIron homeostasis in plants – a brief overview
James M.
Connorton
 *,
Janneke
Balk
*,
Janneke
Balk
 * and
Jorge
Rodríguez-Celma
* and
Jorge
Rodríguez-Celma
 *
*
John Innes Centre and University of East Anglia, Norwich Research Park, Norwich, NR4 7UH, UK. E-mail: james.connorton@jic.ac.uk; janneke.balk@jic.ac.uk; jorge.rodriguez-celma@jic.ac.uk
First published on 28th June 2017
Abstract
Iron plays a crucial role in biochemistry and is an essential micronutrient for plants and humans alike. Although plentiful in the Earth's crust it is not usually found in a form readily accessible for plants to use. They must therefore sense and interact with their environment, and have evolved two different molecular strategies to take up iron in the root. Once inside, iron is complexed with chelators and distributed to sink tissues where it is used predominantly in the production of enzyme cofactors or components of electron transport chains. The processes of iron uptake, distribution and metabolism are overseen by tight regulatory mechanisms, at the transcriptional and post-transcriptional level, to avoid iron concentrations building to toxic excess. Iron is also loaded into seeds, where it is stored in vacuoles or in ferritin. This is important for human nutrition as seeds form the edible parts of many crop species. As such, increasing iron in seeds and other tissues is a major goal for biofortification efforts by both traditional breeding and biotechnological approaches.
Dr Janneke Balk studied Biology at Wageningen University (Netherlands) followed by a DPhil in Biological Sciences at the University of Oxford (UK). In 2005 she received a Royal Society University Research Fellowship to set up her own group at the University of Cambridge and study iron–sulphur cluster assembly in plants. She is currently a Project Leader at the John Innes Centre in Norwich and an Associate Professor at the University of East Anglia. Her research interests are mitochondria, iron–sulphur proteins and iron homeostasis. |
Dr Jorge Rodríguez-Celma studied Biochemistry at the University of Zaragoza (Spain), where he also completed his PhD, which was on proteomics applied to iron deficiency and cadmium toxicity in plants. He moved to the Academia Sinica in Taipei (Taiwan) to do post-doctoral research on iron and manganese-regulated transcription networks. In 2015 he received a Marie Sklodowska Curie Individual Fellowship to study Fe sensing mechanisms in plants at the University of East Anglia and John Innes Centre (UK). |
1 Introduction
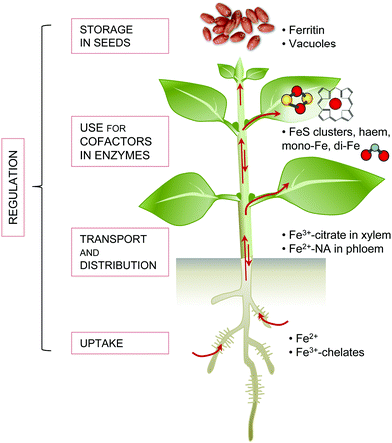
The redox properties of iron make it an essential element for practically all life. Iron is a component of cofactors that carry out electron transfer functions, or facilitate chemical transitions such as hydroxylations, radical-mediated rearrangements and (de)hydration reactions. Iron cofactors also function in oxygen transport, oxygen or iron sensing, or regulation of protein stability. The chloroplasts are particularly rich in iron–sulphur (FeS) proteins such as Photosystem I, ferredoxins and a range of metabolic enzymes. Mitochondria are another hotspot for iron enzymes, such as respiratory complexes containing multiple FeS clusters (complex I and II), a mix of FeS and haem (complex III) or haem and copper (complex IV). The peroxisomes and the endoplasmic reticulum contain haem proteins such as peroxidases and cytochrome P450s, whereas mono- and di-iron enzymes are found in all cell compartments.Iron limitation severely affects plant growth and iron is often a component of agricultural fertilisers used to improve crop yields. Although iron is abundant in the Earth's crust, it is usually present in an oxidised form that is not easily accessible for life. Plants, as primary producers, are the gateway for iron to enter the food chain. Iron-deficiency anaemia is a major human health issue, estimated by the World Health Organization to affect over 30% of the world's population (http://www.who.int/nutrition/topics/ida/en/). On the other hand, excess iron is toxic to cells. Therefore, organisms have evolved intricate mechanisms to take up, metabolise and store iron, and regulate these processes to maintain homeostasis. Fig. 1 gives an overview of the main principles of iron homeostasis in plants. Research over the last decade has identified many key players that deal specifically with iron. These comprise high-affinity uptake transporters and biosynthetic enzymes of organic chelators, transporters for distributing iron to other tissues and storage, enzymes for iron cofactor biosynthesis, plus a regulatory network of transcription factors. Often the finer details of these processes paint a complex picture that can be impenetrable for those new to the field. The aim of this review is to provide an overview of iron homeostasis and how this knowledge can be used to improve human nutrition. We intend it to be an entry-point for those with limited prior knowledge, and hope it may prove useful for teaching purposes. Readers wishing for more detail are referred to the comprehensive review by Kobayashi and Nishizawa (2012),1 or to recent reviews covering specific aspects such as iron uptake and regulation,2 – omics studies applied to Fe deficiency,3 iron mobilisation from soil,4 iron cofactor biosynthesis5 and biofortification.6
2 Iron uptake
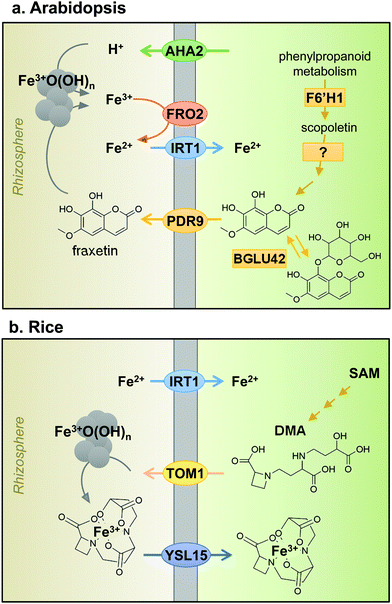
Iron uptake in plants has classically been divided into Strategy I and Strategy II, also known as reducing and chelating strategies, respectively.7 The main difference between both strategies is the oxidation state of iron when taken up by the plant: ferrous Fe2+ for Strategy I and ferric Fe3+ for Strategy II. Iron in the rhizosphere is mainly present as Fe3+ oxyhydrates of low solubility. Tomato and Arabidopsis have served as models for Strategy I (Fig. 2a), in which Fe3+ is reduced by Ferric Reduction Oxidase 2 (FRO2) at the plasma membrane8 before transport across the membrane by Iron-Regulated Transporter 1 (IRT1).9 In addition, plasma-membrane proton pumps such as AHA2 help to acidify the rhizosphere and increase Fe3+ solubility.10 Barley, rice and maize in the grass family (Poaceae) represent Strategy II plants (Fig. 2b), which secrete phytosiderophores, defined as plant-derived small organic molecules with a high affinity for iron.11 Deoxymugineic acid is the most abundant phytosiderophore and is exported by TOM1, the Transporter of Mugineic acid family phytosiderophores, in rice and barley.12 The Fe3+-phytosiderophore chelates are imported by the oligopeptide transporter YS1, first characterised in maize,13 and later in rice (YSL15).14However, the dichotomy of iron uptake into Strategy I and Strategy II plants is perhaps too simplistic when considering recent discoveries in the iron-deficiency response. Strategy I plants were found to export an array of metabolites including organic acids, phenolics, flavonoids and flavins.15 Phenolics were initially hypothesised to help with the solubilisation and reutilisation of apoplastic iron in red clover.16 This feature was not considered part of the iron uptake mechanism until coumarin-derived phenolics were observed in Arabidopsis under high pH conditions.17–19 Coumarins are synthesised using precursors from the phenylpropanoid pathway. The first coumarin in the pathway, scopoletin, is synthesised by the enzyme feruloyl CoA ortho-hydroxylase 1 (F6′H1).17,19,20 While there is some uncertainty about the next intermediate(s) and the precise biosynthetic steps, the active end product is most likely fraxetin.21 An important chemical feature of fraxetin for iron chelation and mobilization is the catechol moiety, two adjacent hydroxyl groups on a benzene ring (Fig. 2a).19,21 The secretion of coumarin chelators is dependent on specific β-glucosidases to remove a glucoside group, such as BGLU42,22 and the ATP-binding cassette transporter PDR9/ABCG37.17,18 Other plant species such as alfalfa (Medicago) and sugar beet secrete flavins instead of coumarins, which also function to facilitate the reduction of ferric iron.17,23 In addition, secretion of the polyamine compound putrescine appears to improve mobilisation of iron inside the plant cell wall.24 Taken together, the results imply that Strategy I plants produce and secrete chelators to the rhizosphere, a characteristic of Strategy II plants. Having said that, mutant studies showed that to take up iron mobilised by coumarins, FRO2 and IRT1 are required, thus IRT1 remains the main route of iron uptake in Arabidopsis.25 Moreover, rice and barley have a functional homologue of IRT1, which mediates the uptake of Fe2+ under low oxygen,26,27 thus blurring the distinction between Strategy I and II even further. Two different uptake strategies for Fe2+ and Fe3+ in the same organism are widespread in Nature. For example, in animals, Fe2+ is transported by the Divalent Metal Transporter DMT1 and Fe3+ is captured by transferrin by the same cells.28 Bacteria take up both Fe2+ and Fe3+-chelates by specific transporters.29
3 Iron distribution and storage
Most iron enters the plant via the root and then needs to be transported to the sink tissues where it is required for iron-dependent enzymes. IRT1 is predominantly localised to the outward facing membrane of epidermal cells30 suggesting that is where iron first enters the symplastic pathway in which cells are connected by plasmodesmata. It is likely that efflux transporters localise to the inner membrane domain of root epidermal cells, but these have not yet been identified.31 NRAMP1 is suggested to cooperate with IRT1 in iron uptake, possible as a low-affinity uptake system.32 Nutrients can also travel through the apoplastic space formed by the cell walls of epidermis and cortex cells to reach the endodermis. Here iron meets a barrier in the form of the Casparian strip, a layer of waterproof lignin, which forces all iron to pass into the symplast. The endodermis can therefore be considered a checkpoint for the translocation of iron into the plant.33 The amount of suberisation of the endodermis changes in response to environmental factors, with iron-deficient plants showing a marked decrease in suberisation that results in an increase in the permeability of the endodermis, allowing more iron to enter the vasculature.34Due to its toxicity and low solubility iron must be complexed to chelators to be translocated effectively without causing damaging redox reactions. In the symplast iron is thought to be transported in the form of Fe2+–nicotianamine (NA) complexes. NA is a non-protein amino acid produced from S-adenosyl methionine by nicotianamine synthase (NAS), encoded by a small gene family in most plant species.35–37 NA is a precursor of mugineic acid (see Section 2) and it also chelates Zn2+ and other divalent cations. Once iron has passed the endodermis, it can be loaded into the xylem for transport to the shoot. This is carried out by the pericycle, a layer of cells inside the endodermis. The xylem comprises dead cells that form a conduit, therefore iron needs to be exported from the symplastic space into the apoplast, possibly by YSL238 and ferroportin,39 although biochemical evidence from transport studies is currently lacking. The dominant form of iron in the xylem is Fe3+–citrate40 and consequently Fe2+ must be oxidised to Fe3+. In addition, citrate efflux is crucial for iron translocation, and this is mediated by the efflux transporter FRD3 in Arabidopsis41 and its orthologue FRDL1 in rice.42
An important sink tissue for iron is the leaves, where it is needed for photosynthesis. Here iron re-enters the symplast and is reduced to Fe2+, mainly by the action of FRO proteins, and is again found as Fe2+–NA. A large proportion of iron is used in the plastids and mitochondria and iron transporters specific for each type of organelle have been identified, see recent reviews.43,44 Iron is remobilised from leaf tissues and reaches other sink organs through the phloem. In Arabidopsis the oligopeptide transporter family protein OPT3 is involved in this process and opt3 mutants have more iron trapped in leaves with less translocated elsewhere such as the seed.45,46
Though present in many tissues, the terminal destination of iron is often considered to be the seed, where iron stores are important during germination before the seedling has developed a root and takes up nutrients from the soil. YSL transporters are involved in seed loading,47 and there is evidence that iron can be delivered to pea embryos as a Fe3+-citrate/malate complex.48
Two major storage mechanisms for iron are proposed: sequestration into vacuoles and into ferritin. The Vacuolar Iron Transporter VIT1 was first identified in Arabidopsis as an orthologue of the yeast iron transporter CCC1. In vit1 mutants, the iron content of embryos is similar to wild type, but the iron no longer accumulates in the vacuoles of the root endodermis and veins.49,50 The action of the efflux transporters NRAMP3 and NRAMP4 releases iron into the cytosol during germination.51 A suppressor screen of nramp3/nramp4 mutants identified mutations in VIT1 that rescue their sensitivity to low iron.52 Genes from the VIT family are also known to be important for iron localisation in rice grains and Brassica seeds.53,54
Ferritins are important iron storage proteins present across the biological kingdoms. Twenty-four subunits form a shell able to store up to a maximum of 4500 Fe3+ ions, although purified plant ferritin, for example from legume seeds, tends to contain approximately 2500 ions.55 The proportion of total iron stored in ferritin in seeds varies among species, with approximately 60% in peas but less than 5% in Arabidopsis seeds.56 In plants, ferritin is predominantly located in the plastids. In cereal grains such as wheat and rice, most iron is present in vacuoles in the aleurone layer which is often removed during grain processing.57 The way in which iron is stored in seeds can affect its bioavailability when consumed, which is of great importance to biofortification studies (see Section 6).
4 Biosynthesis of iron co-factors
The most common forms of iron cofactors are haem, FeS clusters and di-iron centres.5,58 Haem contains one iron atom inserted into an organic tetrapyrrole ring and is non-covalently or covalently (in the case of c-type cytochromes) bound to the protein. In FeS clusters, 2 or 4 iron atoms are bridged by acid-labile sulphides and linked to at least one cysteine sulphur of the protein. The ligands of the iron atom are critical in modifying the precise catalytic properties of an enzyme. For example, all-cysteine FeS clusters occupy the lower range of redox potentials (approximately −300 to +100 mV) and haems the higher range (+50 to +400 mV).58 Di-iron centres, also known as Fe–O–Fe centres, and mono-iron centres are generally bound by histidine, glutamate and aspartate residues and function in hydroxylation and oxygenation reactions, respectively.Because of the toxic nature of free iron, cofactor biosynthesis is a highly controlled process. The biosynthesis pathways of haem and FeS clusters are well characterised (Fig. 3), but little is known about the delivery of iron to these pathways or to mono- and di-iron enzymes. Plants contain relatively little haem in comparison to animals, about 0.5–1% of total iron.59 Haem biosynthesis takes place in the plastids, as a branch of chlorophyll biosynthesis. Both molecules have a tetrapyrrole ring but a different metal (Fe and Mg, respectively) and specific side groups. The starting point for the biosynthesis of tetrapyrrole is glutamate in the form of glutamyl-tRNAGLU, and the pathway involves nine enzymes essentially conserved with bacteria, before insertion of iron by ferrochelatase (FC).60 Land plants express at least two isoforms of FC. Mutant studies in Arabidopsis have shown that FC1 provides haem at all growth stages, whereas FC2 boosts haem levels in chloroplasts during photosynthesis.59,61 Haem is presumably transported from its site of biosynthesis to other cell compartments, but the transporters have not been characterised in plants. In fungi and metazoa, mitochondria are the site of haem biosynthesis, and it has been proposed that plant mitochondria harbour enzyme isoforms for the last two steps, although firm evidence is lacking.
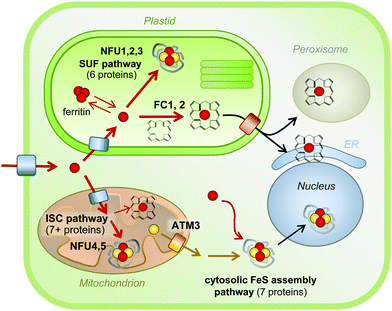
For the biosynthesis of FeS clusters, both plastids and mitochondria harbour complete assembly pathways (see reviews5,62). Cysteine serves as the source of acid-labile sulphur, which is transferred from the active site of the desulphurase enzyme to a scaffold protein where sulphide is combined with iron. The precise mechanistic details of the assembly process remain to be established, but it does require reducing equivalents provided by ferredoxin and/or NADPH.63,64 The plastid and mitochondrial cysteine desulphurases belong to different pathways, called SUF (SUlFur mobilization) and ISC (Iron Sulfur Cluster), respectively. The genes are mostly conserved with those in bacteria. The plastid pathway consists of 6 SUF proteins, all except SUFA essential for plant viability. SUFS and SUFE form the desulphurase activity65 and the SUFB2CD complex is the FeS cluster scaffold.66 SUFA and 3 partially redundant NFU proteins are involved in carrying FeS clusters from the scaffold to recipient FeS proteins. In the mitochondrial ISC pathway, a similar division in desulphurase activity (NFS1 and ISD11), scaffold (ISU) and carrier proteins (NFU4 and NFU5) can be made. FeS clusters cannot cross membranes – except as part of folded proteins through the twin-arginine pathway – and a separate set of at least 7 proteins for Cytosolic Iron–sulphur protein Assembly (CIA) are required for the activity of FeS enzymes in the cytosol and nucleus.67 The CIA pathway is functionally connected to the mitochondria, which are thought to provide sulphur exported by an ATP Binding Cassette transporter.68,69 Interestingly, forward and reverse genetics studies have highlighted the importance of Arabidopsis CIA proteins in plant development, because of the role of FeS enzymes in plant hormone biosynthesis, DNA demethylation and DNA repair.70–74
5 Regulation of iron homeostasis
An efficient regulatory system that senses the iron status of the plant, and adjusts homeostasis accordingly, is crucial for ensuring enough iron reaches the tissues where it is needed without accumulating to toxic levels. Plants adapt their root morphology to iron-limiting conditions by increasing the density of root hairs and the number of lateral roots. The greater surface area extends contact between the epidermis and the rhizosphere, and the lateral roots help to explore fresh soil.75 These macroscopic changes have been studied mostly from the perspective of plant development.76 The entry of iron into the symplast is controlled at the endodermis (see Section 3). The precise molecular mechanisms that link iron availability to changes in root morphology or permeability are not yet known.Over the past 15 years, great progress has been made in identifying a large number of transcriptional regulators of iron homeostasis. However, this has led to a somewhat bewildering network of basic Helix-Loop-Helix (bHLH) transcription factors that regulate the iron deficiency response (see Table 1 and Fig. 4). The first transcription factor that was cloned was FER from tomato,77 followed shortly by its functional orthologue in Arabidopsis called FER-like Iron deficiency-induced Transcription factor (FIT). FIT is expressed in roots only and required for up-regulating iron uptake genes such as FRO2 and IRT1.78–80 FIT cannot act alone but must form a heterodimer with one of four other bHLH proteins, bHLH38, bHLH39, bHLH100 and bHLH101, to bind to the promoters of IRT1 and FRO2, as was elegantly shown in yeast.81,82 Mutant studies in Arabidopsis have shown that the four partner proteins are partially redundant,82 but they may fine-tune the response by activating different downstream genes.83
| Gene | Expression pattern | Downstream genes | Ref. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| a Rice genes. The closest homologue of IRO2 is bHLH39 in Arabidopsis. For IRO3, the closest homologue is PYE. There are no homologues of IDEF1 and IDEF2 in Arabidopsis. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FIT1 | Root | Direct: FRO2, IRT1 | 78–81 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Other: MYB10, MYB72, F6'H1, many other genes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Subgroup Ib: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| bHLH38, bHLH39 | All tissues | FRO2, IRT1, | 81–83 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| bHLH100, bHLH101 | Phenylalanine metabolism | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Subgroup IVc: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| bHLH034 | Vasculature | bHLH38/39/100/101, PYE | 85 and 86 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| bHLH104 | Vasculature | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| bHLH105 (ILR3), bHLH115 | Vasculature, meristems | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MYB10 | Roots | NAS4 | 22 and 87 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MYB72 | Coumarin biosynthesis, BGLU42 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| WRKY46 | Roots | Direct: VITL1 | 88 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NAS2? | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PYE | Roots and shoots | Direct: NAS4, ZIF1, FRO3 | 84 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IDEF1 | Roots (phloem) and leaves (mesophyll) | Phytosiderophore biosynthesis, OsIRT1, OsIRO2 | 92 and 123 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IDEF2 | Roots and leaves (vasculature) | YSL2 | 92 and 124 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IRO2 | Roots and shoots | Phytosiderophore biosynthesis, YSL15 | 91 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IRO3 | Roots and shoots | NAS1, NAS2, IRO2 | 93 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
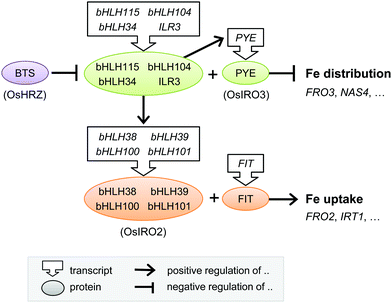 | ||
| Fig. 4 Transcriptional regulation of the iron deficiency response in plants. Diagram depicting the core transcriptional regulators and their functional relationship. The rice homologues are given in parentheses, but note that downstream gene targets in rice may differ. Further details can be found in the main text and in Table 1. bHLH, basic Helix-Loop-Helix protein; BTS, BRUTUS; FIT, FER-like Iron deficiency-induced Transcription factor; PYE, POPEYE. | ||
A cell-type specific microarray study of iron-deficient Arabidopsis roots identified another bHLH protein POPEYE (PYE) which is part of a regulatory network independent of FIT.84 PYE interacts with two bHLH transcription factors, ILR3 (bHLH105) and bHLH115, and is itself regulated by dimers of ILR3 and bHLH104, or ILR3 and bHLH34.85,86 Other families of transcription factors have also been implicated in the iron deficiency response. MYB10 and MYB72 of the MYB family regulate expression of NAS487 and the production of coumarins.22 Recently, a member of the WRKY family, WRKY46, was demonstrated to regulate expression of NAS2 and the VIT-Like 1 gene.88
There is less information about the transcriptional networks governing iron uptake in monocotyledonous plants. Studies have focused on finding transcription factors that bind to the iron-responsive motifs in the promoter of Iron Deficiency Specific clone 2 (IDS2).89 This strategy led to the discovery of transcription factors IDEF1 and IDEF2 in rice,90 which control the expression of phytosiderophore biosynthesis and YSL2.92 Part of the transcriptional regulatory networks found in Arabidopsis, including members of the bHLH transcription family, are conserved in rice. IRO2 in rice is a close homologue of bHLH39 and positively regulates phytosiderophore biosynthesis and YSL15.91 IRO3 is the rice orthologue of PYE and negatively controls the transcript levels of IRO2 and NAS.93
The iron-deficiency response results in an increase in iron uptake which could inadvertently lead to overload if iron becomes suddenly available in the environment, such as after rainfall. Several post-transcriptional mechanisms have been observed that rapidly stop iron uptake. For example, IRT1 is continuously recycled from the plasma membrane via ubiquination and internalisation, and an E3 ligase responsible for its ubiquination has been identified.94 In addition to swiftly degrading the uptake machinery, it is also necessary to stop the transcriptional response. FIT has been reported to be actively turned over in response to ethylene and nitric oxide,83,95,96 but no E3 ligase responsible for FIT degradation has been found to date. Ultimately, all these mechanisms must relate to the iron status in the cell, assuming the existence of Fe-binding regulators such as Fur in bacteria, Aft1 in yeast, or IRPs and FBXL5 in mammals. Recent studies in plants suggest that a small family of hemerythrin E3 ligases, including BRUTUS (BTS) in Arabidopsis97 and HRZ in rice,98 may sense iron and act as negative regulators of the iron deficiency response. The three hemerythrin domains in the N-terminus of BTS and HRZ have conserved His-xxx-Glu motifs likely to bind a di-iron centre. The C-terminal domain has 45% homology to plant and mammalian E3 ligases that target transcription factors for ubiquitination and subsequent turnover. Interestingly, BTS was found to interact with selected bHLH proteins,97 but many questions remain. Such as how the hemerythrin domains sense intracellular Fe levels, why there are three hemerythrin domains, and how iron binding modulates the E3 ligase activity of the C-terminal domain.
6 Biofortification of crops with iron
As our understanding of the mechanisms of iron uptake, transport and homeostasis increase, more applications of this knowledge are being explored to biofortify crops. The five most widely-consumed crops in the developing world are maize, rice, wheat, pulses and cassava (http://www.fao.org/in-action/inpho/crop-compendium/en/), and as such biofortification efforts have focussed on these species. As mentioned previously, plants exhibit tight homeostatic control to prevent accumulation of iron where it is not needed, and this may limit iron redistribution to edible tissues such as seeds. Any successful biofortification strategy must bypass these mechanisms without causing physiological damage to the plant. Progress is being made through two main strategies: traditional breeding and modern technology including transgenics (Table 2).| Crop | Details | Strategy employed | Fe content (fold increase over control) | Ref. |
|---|---|---|---|---|
| Beans | High iron landrace | Traditional breeding for high iron content – molecular details unknown | 1.7× in whole beans | 102 and 105 |
| Cassava | AtVIT1 with PATATIN promoter | Transgenic: increased iron storage in target tissue | 3–4× in storage roots | 107 |
| Rice | Combination of overexpressing AtNAS1, OsNAS2, GmFERRITIN and AfPHYTASE | Transgenic: increased iron translocation and storage, degradation of phytate | 6× in T4 polished seeds | 125 |
| Wheat | OsNAS2 with ZmUBIQUITIN promoter | Transgenic: increased translocation. Co-transformation with ferritin had no synergistic effect | 2.5× in whole grains | 126 |
Traditional breeding methods have existed for thousands of years and given rise to many useful crop varieties. However, breeders have concentrated primarily on increasing yield, and as a consequence the levels of iron have been diluted by increased starch.99 One current drive in crop research is to restore old traits in modern varieties to make them more nutritious. For example, the NAM-B1 transcription factor has been lost from modern wheat but is present in older varieties where it advances senescence and leads to higher iron, zinc and protein content in the grain,100 a discovery which has since informed breeding programmes.101 In a separate approach, taking advantage of natural variation has made it possible to breed for higher iron levels in crops such as beans and pearl millet.102,103 In pilot studies, in part funded by HarvestPlus (http://www.harvestplus.org), high iron varieties of these crops have been used successfully to improve iron status in iron-deficient and anaemic women and children in Rwanda and India.103–105
Alternative approaches for iron biofortification use transgenic and cisgenic technologies. The advantage of these practices over traditional breeding is that it is possible to target genes of interest directly, altering expression of endogenous genes (cisgenics) or introducing genetic material from other species (transgenics). Such approaches have successfully biofortified rice by constitutively expressing endogenous NAS genes,106 and cassava by expressing Arabidopsis VIT1 in storage roots.107 Iron storage in both ferritin and vacuoles have been targeted by cisgenic strategies to biofortify wheat grain.108,109 Increasing VIT expression in the wheat endosperm redirects iron to this part of the grain (Fig. 5). A concern about using transgenics in biofortification is that, mainly due to low substrate specificity of the IRT1 transporter, levels of toxic metals such as cadmium may increase alongside iron.110 This has so far not been a problem, for example overexpression of barley Yellow Stripe 1 (HvYS1) led specifically to increases in iron.111
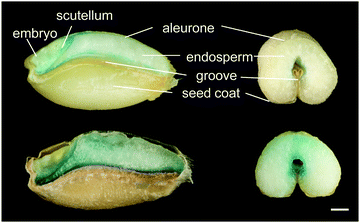 | ||
| Fig. 5 Perls’ Prussian Blue staining for iron in wheat grains from control line (top) and high-iron line expressing TaVIT2 in the endosperm (bottom). Grains were dissected longitudinally (left) or transversely (right) using a platinum-coated blade. For further details see Connorton et al., 2017.109 Scale bar = 1 mm. | ||
One limitation is that only relatively few crop varieties can be transformed using current techniques, and so traits developed in “transformable” varieties must be introduced into elite varieties by potentially time-consuming crosses. Modern genetic technologies such as TILLING and CRISPR/Cas are not classed as transgenic, and could prove valuable in producing iron biofortified crops. TILLING populations have been produced in species such as wheat, which allow researchers to knock out the function of specific genes.112 As well as being a useful tool for studying gene function, this also unlocks the potential to suppress the function of negative regulators of iron accumulation in specific tissues.
A further obstacle when biofortifying cereals is that iron and other minerals are often poorly bioavailable. The main reason for this is the presence of anti-nutrient compounds such as phytate (myo-inositol-1,2,3,4,5,6-hexakisphosphate) and polyphenols which chelate minerals and prevent them being absorbed by the gut.113 Phytate is a phosphate storage compound abundant in the aleurone and seed coat of cereal grains. Several strategies have been explored to decrease the level of phytate in crops, such as breeding programs aided by identification of relevant Quantitative Trait Loci (QTL)114 and expression of phytase genes.115 Recently published research into phosphate transport showed that rice mutants lacking the SPDT phosphate transporter had a sharp decrease in grain phytate levels as well as a modest increase in iron and other minerals.116 It should also be noted that the way plant material is processed post-harvest can affect bioavailability: iron in white flour is generally more bioavailable than wholemeal flour,117 sourdough bread is produced with bacteria that have naturally occurring phytases,118 and the micromilling of flour can aid bioavailability of minerals.119 Future biofortification strategies would take account of all these factors, combining high iron in edible parts along with low phytate, and using post-harvest processes that increase bioavailability. Current research efforts tend to focus on one of these factors at a time, although more combinatorial studies are emerging (Table 2).
7 Conclusions and future perspectives
Great progress has been made over the past decade in understanding the mechanisms of iron homeostasis in plants. More studies are emerging where information gathered in model plants is being used to study crops, and to generate varieties with higher levels of iron. Many topics do remain to be addressed, however, which will influence future research directions. For example, the substrate specificities of the many transporters involved in iron transport – from cell-to-cell and between intracellular compartments – need to be demonstrated by biochemical studies. Long-distance signalling between shoots and roots has not been discussed in this review, because the components are unknown. The transcriptional networks involved in iron homeostasis have rapidly expanded, but many redundancies between gene functions have been reported and a unified model of the signalling cascade is lacking. cis-Element prediction tools have recently been refined,120–122 but approaches such as ChIP-seq analysis would more clearly define transcriptional regulons and help establish the hierarchy of the different transcription factors. In addition, a tissue- or cell-specific view of iron homeostasis would be insightful, and the function of the cell wall is usually overlooked. A major unresolved question is how iron in plants is sensed, what the precise role of the BTS/HRZ proteins is, and how the iron status is signalled to permeability changes of the endodermis. There is also still limited information available on how the speciation of iron impacts on the bioavailability of iron in plant foods. Overall, these are exciting times, with traditional physiological and genetic studies of iron homeostasis being enhanced by genomics and metabolomics approaches.Acknowledgements
We are grateful for funding from HarvestPlus (J. M. C.) and the BBSRC BB/N001079/1, BB/J004561/1. J. R.-C. was funded by a Marie Sklodowska Curie fellowship from the EU H2020 programme.References
- T. Kobayashi and N. K. Nishizawa, Iron uptake, translocation, and regulation in higher plants, Annu. Rev. Plant Biol., 2012, 63, 131–152 CrossRef CAS PubMed.
- T. Brumbarova, P. Bauer and R. Ivanov, Molecular mechanisms governing Arabidopsis iron uptake, Trends Plant Sci., 2015, 20, 124–133 CrossRef CAS PubMed.
- W. Li and P. Lan, The understanding of the plant iron deficiency responses in Strategy I plants and the role of ethylene in this process by omic approaches, Front. Plant Sci., 2017, 8, 40 Search PubMed.
- H. H. Tsai and W. Schmidt, Mobilization of iron by plant-borne coumarins, Trends Plant Sci., 2017, 22, 538–548 CrossRef CAS PubMed.
- J. Balk and T. A. Schaedler, Iron cofactor assembly in plants, Annu. Rev. Plant Biol., 2014, 65, 125–153 CrossRef CAS PubMed.
- M. W. Vasconcelos, W. Gruissem and N. K. Bhullar, Iron biofortification in the 21st century: setting realistic targets, overcoming obstacles, and new strategies for healthy nutrition, Curr. Opin. Biotechnol., 2017, 44, 8–15 CrossRef CAS PubMed.
- V. Römheld and H. Marschner, Evidence for a specific uptake system for iron phytosiderophores in roots of grasses, Plant Physiol., 1986, 80, 175–180 CrossRef.
- N. J. Robinson, C. M. Procter, E. L. Connolly and M. L. Guerinot, A ferric-chelate reductase for iron uptake from soils, Nature, 1999, 397, 694–697 CrossRef CAS PubMed.
- D. Eide, M. Broderius, J. Fett and M. L. Guerinot, A novel iron-regulated metal transporter from plants identified by functional expression in yeast, Proc. Natl. Acad. Sci. U. S. A., 1996, 93, 5624–5628 CrossRef CAS.
- S. Santi and W. Schmidt, Dissecting iron deficiency-induced proton extrusion in Arabidopsis roots, New Phytol., 2009, 183, 1072–1084 CrossRef CAS PubMed.
- K. Higuchi, et al., Cloning of nicotianamine synthase genes, novel genes involved in the biosynthesis of phytosiderophores, Plant Physiol., 1999, 119, 471–480 CrossRef CAS PubMed.
- T. Nozoye, et al., Phytosiderophore efflux transporters are crucial for iron acquisition in graminaceous plants, J. Biol. Chem., 2011, 286, 5446–5454 CrossRef CAS PubMed.
- C. Curie, et al., Maize yellow stripe1 encodes a membrane protein directly involved in Fe(III) uptake, Nature, 2001, 409, 346–349 CrossRef CAS PubMed.
- H. Inoue, et al., Rice OsYSL15 is an iron-regulated iron(III)-deoxymugineic acid transporter expressed in the roots and is essential for iron uptake in early growth of the seedlings, J. Biol. Chem., 2009, 284, 3470–3479 CrossRef CAS PubMed.
- S. Cesco, G. Neumann, N. Tomasi, R. Pinton and L. Weisskopf, Release of plant-borne flavonoids into the rhizosphere and their role in plant nutrition, Plant Soil, 2010, 329, 1–25 CrossRef CAS.
- C. W. Jin, et al., Iron deficiency-induced secretion of phenolics facilitates the reutilization of root apoplastic iron in red clover, Plant Physiol., 2007, 144, 278–285 CrossRef CAS PubMed.
- J. Rodríguez-Celma, et al., Mutually exclusive alterations in secondary metabolism are critical for the uptake of insoluble iron compounds by Arabidopsis and Medicago truncatula, Plant Physiol., 2013, 162, 1473–1485 CrossRef PubMed.
- P. Fourcroy, et al., Involvement of the ABCG37 transporter in secretion of scopoletin and derivatives by Arabidopsis roots in response to iron deficiency, New Phytol., 2014, 201, 155–167 CrossRef CAS PubMed.
- N. B. Schmid, et al., Feruloyl-CoA 6′-Hydroxylase1-dependent coumarins mediate iron acquisition from alkaline substrates in Arabidopsis, Plant Physiol., 2014, 164, 160–172 CrossRef CAS PubMed.
- K. Kai, et al., Scopoletin is biosynthesized via ortho-hydroxylation of feruloyl CoA by a 2-oxoglutarate-dependent dioxygenase in Arabidopsis thaliana, Plant J., 2008, 55, 989–999 CrossRef CAS PubMed.
- P. Sisó-Terraza, et al., Accumulation and secretion of coumarinolignans and other coumarins in Arabidopsis thaliana roots in response to iron deficiency at high pH, Front. Plant Sci., 2016, 7, 1711 Search PubMed.
- C. Zamioudis, J. Hanson and C. M. J. Pieterse, β-Glucosidase BGLU42 is a MYB72-dependent key regulator of rhizobacteria-induced systemic resistance and modulates iron deficiency responses in Arabidopsis roots, J. Physiol., 2014, 204, 368–379 CAS.
- P. Sisó-Terraza, J. J. Rios, J. Abadía, A. Abadía and A. Álvarez-Fernández, Flavins secreted by roots of iron-deficient Beta vulgaris enable mining of ferric oxide via reductive mechanisms, New Phytol., 2016, 209, 733–745 CrossRef PubMed.
- X. F. Zhu, B. Wang, W. F. Song, S. J. Zheng and R. F. Shen, Putrescine alleviates iron deficiency via NO-dependent reutilization of root cell-wall Fe in Arabidopsis, Plant Physiol., 2016, 170, 558–567 CrossRef CAS PubMed.
- P. Fourcroy, N. Tissot, F. Gaymard, J. F. Briat and C. Dubos, Facilitated Fe nutrition by phenolic compounds excreted by the Arabidopsis ABCG37/PDR9 transporter requires the IRT1/FRO2 high-affinity root Fe2+ transport system, Mol. Plant, 2016, 9, 485–488 CrossRef CAS PubMed.
- Y. Ishimaru, et al., Rice plants take up iron as an Fe3+-phytosiderophore and as Fe2, Plant J., 2006, 45, 335–346 CrossRef CAS PubMed.
- P. Pedas, et al., Manganese efficiency in barley: identification and characterization of the metal ion transporter HvIRT1, Plant Physiol., 2008, 148, 455–466 CrossRef CAS PubMed.
- D. J. R. Lane, et al., Cellular iron uptake, trafficking and metabolism: Key molecules and mechanisms and their roles in disease, Biochim. Biophys. Acta, Mol. Cell Res., 2015, 1853, 1130–1144 CrossRef CAS PubMed.
- L. D. Palmer and E. P. Skaar, Transition metals and virulence in bacteria, Annu. Rev. Genet., 2016, 50, 67–91 CrossRef CAS PubMed.
- M. Barberon, et al., Polarization of IRON-REGULATED TRANSPORTER 1 (IRT1) to the plant-soil interface plays crucial role in metal homeostasis, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 8293–8298 CrossRef CAS PubMed.
- G. Dubeaux, E. Zelazny and G. Vert, Getting to the root of plant iron uptake and cell-cell transport: polarity matters!, Commun. Integr. Biol., 2015, 8, e1038441 CrossRef PubMed.
- L. Castaings, et al., The high-affinity metal transporters NRAMP1 and IRT1 team up to take up iron under sufficient metal provision, Sci. Rep., 2016, 6, 37222 CrossRef CAS PubMed.
- M. Barberon, The endodermis as a checkpoint for nutrients, New Phytol., 2017, 213, 1604–1610 CrossRef CAS PubMed.
- M. Barberon, et al., Adaptation of root function by nutrient-induced plasticity of endodermal differentiation, Cell, 2016, 164, 447–459 CrossRef CAS PubMed.
- H. Inoue, et al., Three rice nicotianamine synthase genes, OsNAS1, OsNAS2 and OsNAS3 are expressed in cells involved in long-distance transport of iron and differentially regulated by iron, Plant J., 2003, 36, 366–381 CrossRef CAS PubMed.
- M. Klatte, et al., The analysis of Arabidopsis nicotianamine synthase mutants reveals functions for nicotianamine in seed iron loading and iron deficiency responses, Plant Physiol., 2009, 150, 257–271 CrossRef CAS PubMed.
- J. Bonneau, U. Baumann, J. Beasley, Y. Li and A. A. T. Johnson, Identification and molecular characterization of the nicotianamine synthase gene family in bread wheat, Plant Biotechnol. J., 2016, 14, 2228–2239 CrossRef CAS PubMed.
- R. J. DiDonato, L. A. Roberts, T. Sanderson, R. B. Eisley and E. L. Walker, Arabidopsis Yellow Stripe-Like2 (YSL2): A metal-regulated gene encoding a plasma membrane transporter of nicotianamine-metal complexes, Plant J., 2004, 39, 403–414 CrossRef CAS PubMed.
- J. Morrissey, et al., The ferroportin metal efflux proteins function in iron and cobalt homeostasis in Arabidopsis, Plant Cell, 2009, 21, 3326–3338 CrossRef CAS PubMed.
- R. Rellán-Alvarez, et al., Identification of a tri-iron(III), tri-citrate complex in the xylem sap of iron-deficient tomato resupplied with iron: new insights into plant iron long-distance transport, Plant Cell Physiol., 2010, 51, 91–102 CrossRef PubMed.
- L. S. Green and E. E. Rogers, FRD3 controls iron localization in Arabidopsis, Plant Physiol., 2004, 136, 2523–2531 CrossRef CAS PubMed.
- K. Yokosho, N. Yamaji and J. F. Ma, OsFRDL1 expressed in nodes is required for distribution of iron to grains in rice, J. Exp. Bot., 2016, 67, 5485–5494 CrossRef CAS PubMed.
- G. Finazzi, et al., Ions channels/transporters and chloroplast regulation, Cell Calcium, 2015, 58, 86–97 CrossRef CAS PubMed.
- K. Bashir, S. Rasheed, T. Kobayashi, M. Seki and N. K. Nishizawa, Regulating subcellular metal homeostasis: the key to crop improvement, Front. Plant Sci., 2016, 7, 1192 Search PubMed.
- Z. Zhai, et al., OPT3 is a phloem-specific iron transporter that is essential for systemic iron signaling and redistribution of iron and cadmium in Arabidopsis, Plant Cell, 2014, 26, 2249–2264 CrossRef CAS PubMed.
- D. G. Mendoza-Cózatl, et al., OPT3 is a component of the iron-signaling network between leaves and roots and misregulation of OPT3 leads to an over-accumulation of cadmium in seeds, Mol. Plant, 2014, 7, 1455–1469 CrossRef PubMed.
- M. Le Jean, A. Schikora, S. Mari, J.-F. Briat and C. Curie, A loss-of-function mutation in AtYSL1 reveals its role in iron and nicotianamine seed loading, Plant J., 2005, 44, 769–782 CrossRef CAS PubMed.
- L. Grillet, et al., Ascorbate efflux as a new strategy for iron reduction and transport in plants, J. Biol. Chem., 2014, 289, 2515–2525 CrossRef CAS PubMed.
- S. A. Kim, et al., Localization of iron in Arabidopsis seed requires the vacuolar membrane transporter VIT1, Science, 2006, 314, 1295–1298 CrossRef CAS PubMed.
- H. Roschzttardtz, G. Conéjéro, C. Curie and S. Mari, Identification of the endodermal vacuole as the iron storage compartment in the Arabidopsis embryo, Plant Physiol., 2009, 151, 1329–1338 CrossRef CAS PubMed.
- V. Lanquar, et al., Mobilization of vacuolar iron by AtNRAMP3 and AtNRAMP4 is essential for seed germination on low iron, EMBO J., 2005, 24, 4041–4051 CrossRef CAS PubMed.
- V. Mary, et al., Bypassing iron storage in endodermal vacuoles rescues the iron mobilization defect in the nramp3 nramp4 double mutant, Plant Physiol., 2015, 169, 748–759 CrossRef CAS PubMed.
- Y. Zhang, Y.-H. Xu, H.-Y. Yi and J.-M. Gong, Vacuolar membrane transporters OsVIT1 and OsVIT2 modulate iron translocation between flag leaves and seeds in rice, Plant J., 2012, 72, 400–410 CrossRef CAS PubMed.
- W. Zhu, et al., Vacuolar iron transporter BnMEB2 is involved in enhancing iron tolerance of Brassica napus, Front. Plant Sci., 2016, 7, 1353 Search PubMed.
- E. C. Theil, Ferritin protein nanocages use ion channels, catalytic sites, and nucleation channels to manage iron/oxygen chemistry, Curr. Opin. Chem. Biol., 2011, 15, 304–311 CrossRef CAS PubMed.
- M. Zielińska-Dawidziak, Plant ferritin—A source of iron to prevent its deficiency, Nutrients, 2015, 7, 1184–1201 CrossRef PubMed.
- B. Kyriacou, et al., Localization of iron in rice grain using synchrotron X-ray fluorescence microscopy and high resolution secondary ion mass spectrometry, J. Cereal Sci., 2014, 59, 173–180 CrossRef CAS.
- J. J. R. Fraústo da Silva and R. J. P. Williams, The biological chemistry of the elements: The inorganic chemistry of life, Oxford University Press, 2001 Search PubMed.
- N. A. Espinas, et al., Allocation of heme is differentially regulated by ferrochelatase isoforms in Arabidopsis cells, Front. Plant Sci., 2016, 7, 1326 Search PubMed.
- P. Brzezowski, A. S. Richter and B. Grimm, Regulation and function of tetrapyrrole biosynthesis in plants and algae, Biochim. Biophys. Acta, 2015, 1847, 968–985 CrossRef CAS PubMed.
- M. Scharfenberg, et al., Functional characterization of the two ferrochelatases in Arabidopsis thaliana, Plant, Cell Environ., 2015, 38, 280–298 CrossRef CAS PubMed.
- J. Couturier, B. Touraine, J.-F. Briat, F. Gaymard and N. Rouhier, The iron-sulfur cluster assembly machineries in plants: current knowledge and open questions, Front. Plant Sci., 2013, 4, 259 Search PubMed.
- A. Picciocchi, R. Douce and C. Alban, The plant biotin synthase reaction: Identification and characterization of essential mitochondrial accessory protein components, J. Biol. Chem., 2003, 278, 24966–24975 CrossRef CAS PubMed.
- Y. Takahashi, A. Mitsui, Y. Fujita and H. Matsubara, Roles of ATP and NADPH in formation of the Fe-S cluster of spinach ferredoxin, Plant Physiol., 1991, 95, 104–110 CrossRef CAS PubMed.
- D. Van Hoewyk, et al., Chloroplast iron-sulfur cluster protein maturation requires the essential cysteine desulfurase CpNifS, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 5686–5691 CrossRef PubMed.
- X. Hu, Y. Kato, A. Sumida, A. Tanaka and R. Tanaka, The SUFBC 2 D complex is required for the biogenesis of all major classes of plastid Fe-S proteins, Plant J., 2017, 90, 235–248 CrossRef CAS PubMed.
- D. J. A. Netz, J. Mascarenhas, O. Stehling, A. J. Pierik and R. Lill, Maturation of cytosolic and nuclear iron-sulfur proteins, Trends Cell Biol., 2013, 24, 303–312 CrossRef PubMed.
- D. G. Bernard, Y. Cheng, Y. Zhao and J. Balk, An allelic mutant series of ATM3 reveals its key role in the biogenesis of cytosolic iron-sulfur proteins in Arabidopsis, Plant Physiol., 2009, 151, 590–602 CrossRef CAS PubMed.
- T. A. Schaedler, et al., A conserved mitochondrial ATP-binding cassette transporter exports glutathione polysulfide for cytosolic metal cofactor assembly, J. Biol. Chem., 2014, 289, 23264–23274 CrossRef CAS PubMed.
- D. Buzas, M. Nakamura and T. Kinoshita, Epigenetic role for the conserved Fe-S cluster biogenesis protein AtDRE2 in Arabidopsis thaliana, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 13565–13570 CrossRef CAS PubMed.
- X. Wang, et al., The cytosolic Fe-S cluster assembly component MET18 is required for the full enzymatic activity of ROS1 in active DNA demethylation, Sci. Rep., 2016, 6, 26443 CrossRef CAS PubMed.
- C. G. Duan, et al., MET18 connects the cytosolic iron-sulfur cluster assembly pathway to active DNA demethylation in Arabidopsis, PLoS Genet., 2015, 11, 1–19 Search PubMed.
- Y. F. Han, et al., The cytosolic iron-sulfur cluster assembly protein MMS19 regulates transcriptional gene silencing, DNA repair, and flowering time in Arabidopsis, PLoS One, 2015, 10, 1–21 Search PubMed.
- E. L. Bastow, K. Bych, J. C. Crack, N. E. Le Brun and J. Balk, NBP35 interacts with DRE2 in the maturation of cytosolic iron-sulfur proteins in Arabidopsis thaliana, Plant J., 2017, 89, 590–600 CrossRef CAS PubMed.
- G. Li, H. J. Kronzucker and W. Shi, The response of the root apex in plant adaptation to iron heterogeneity in soil, Front. Plant Sci., 2016, 7, 1–7 Search PubMed.
- J. Mora-Macías, et al., Malate-dependent Fe accumulation is a critical checkpoint in the root developmental response to low phosphate, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, E3563–E3572 CrossRef PubMed.
- H.-Q. Ling, P. Bauer, Z. Bereczky, B. Keller and M. Ganal, The tomato fer gene encoding a bHLH protein controls iron-uptake responses in roots, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 13938–13943 CrossRef CAS PubMed.
- E. P. Colangelo and M. L. Guerinot, The essential basic Helix-Loop-Helix protein FIT1 is required for the iron deficiency response, Plant Cell, 2004, 16, 3400–3412 CrossRef CAS PubMed.
- M. Jakoby, H. Y. Wang, W. Reidt, B. Weisshaar and P. Bauer, FRU (BHLH029) is required for induction of iron mobilization genes in Arabidopsis thaliana, FEBS Lett., 2004, 577, 528–534 CrossRef CAS PubMed.
- Y. X. Yuan, J. Zhang, D. W. Wang and H. Q. Ling, AtbHLH29 of Arabidopsis thaliana is a functional ortholog of tomato FER involved in controlling iron acquisition in strategy I plants, Cell Res., 2005, 15, 613–621 CrossRef CAS PubMed.
- Y. Yuan, et al., FIT interacts with AtbHLH38 and AtbHLH39 in regulating iron uptake gene expression for iron homeostasis in Arabidopsis, Cell Res., 2008, 18, 385–397 CrossRef CAS PubMed.
- N. Wang, et al., Requirement and functional redundancy of Ib subgroup bHLH proteins for iron deficiency responses and uptake in Arabidopsis thaliana, Mol. Plant, 2013, 6, 503–513 CrossRef CAS PubMed.
- A. B. Sivitz, V. Hermand, C. Curie and G. Vert, Arabidopsis bHLH100 and bHLH101 control iron homeostasis via a FIT-independent pathway, PLoS One, 2012, 7, e44843 CAS.
- T. A. Long, et al., The bHLH transcription factor POPEYE regulates response to iron deficiency in Arabidopsis roots, Plant Cell, 2010, 22, 2219–2236 CrossRef CAS PubMed.
- J. Zhang, et al., The bHLH transcription factor bHLH104 interacts with IAA-LEUCINE RESISTANT3 and modulates iron homeostasis in Arabidopsis, Plant Cell, 2015, 27, 787–805 CrossRef CAS PubMed.
- X. Li, H. Zhang, Q. Ai, G. Liang and D. Yu, Two bHLH transcription factors, bHLH34 and bHLH104, regulate iron homeostasis in Arabidopsis thaliana, Plant Physiol., 2016, 170, 2478–2493 CrossRef CAS PubMed.
- C. M. Palmer, M. N. Hindt, H. Schmidt, S. Clemens and M. L. Guerinot, MYB10 and MYB72 are required for growth under iron-limiting conditions, PLoS Genet., 2013, 9, e1003953 Search PubMed.
- J. Y. Yan, et al., A WRKY transcription factor regulates Fe translocation under Fe deficiency, Plant Physiol., 2016, 171, 2017–2027 CrossRef PubMed.
- T. Kobayashi, et al., Identification of novel cis-acting elements, IDE1 and IDE2, of the barley IDS2 gene promoter conferring iron-deficiency-inducible, root-specific expression in heterogeneous tobacco plants, Plant J., 2003, 36, 780–793 CrossRef CAS PubMed.
- T. Kobayashi, et al., The transcription factor IDEF1 regulates the response to and tolerance of iron deficiency in plants, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 19150–19155 CrossRef CAS PubMed.
- Y. Ogo, et al., The rice bHLH protein OsIRO2 is an essential regulator of the genes involved in Fe uptake under Fe-deficient conditions, Plant J., 2007, 51, 366–377 CrossRef CAS PubMed.
- T. Kobayashi, et al., The spatial expression and regulation of transcription factors IDEF1 and IDEF2, Ann. Bot., 2010, 105, 1109–1117 CrossRef CAS PubMed.
- L. Zheng, et al., Identification of a novel iron regulated basic helix-loop-helix protein involved in Fe homeostasis in Oryza sativa, BMC Plant Biol., 2010, 10, 166 CrossRef PubMed.
- L.-J. Shin, et al., IRT1 DEGRADATION FACTOR1, a RING E3 ubiquitin ligase, regulates the degradation of IRON-REGULATED TRANSPORTER1 in Arabidopsis, Plant Cell, 2013, 25, 3039–3051 CrossRef CAS PubMed.
- S. Lingam, et al., Interaction between the bHLH transcription factor FIT and ETHYLENE INSENSITIVE3/ETHYLENE INSENSITIVE3-LIKE1 reveals molecular linkage between the regulation of iron acquisition and ethylene signaling in Arabidopsis, Plant Cell, 2011, 23, 1815–1829 CrossRef CAS PubMed.
- J. Meiser, S. Lingam and P. Bauer, Posttranslational regulation of the iron deficiency Basic Helix-Loop-Helix transcription factor FIT is affected by iron and nitric oxide, Plant Physiol., 2011, 157, 2154–2166 CrossRef CAS PubMed.
- D. Selote, R. Samira, A. Matthiadis, J. W. Gillikin and T. A. Long, Iron-binding E3 ligase mediates iron response in plants by targeting basic Helix-Loop-Helix transcription factors, Plant Physiol., 2015, 167, 273–286 CrossRef CAS PubMed.
- T. Kobayashi, et al., Iron-binding haemerythrin RING ubiquitin ligases regulate plant iron responses and accumulation, Nat. Commun., 2013, 4, 2792 Search PubMed.
- M. S. Fan, et al., Evidence of decreasing mineral density in wheat grain over the last 160 years, J. Trace Elem. Med. Biol., 2008, 22, 315–324 CAS.
- C. Uauy, A. Distelfeld, T. Fahima, A. Blechl and J. A. Dubcovsky, NAC Gene regulating senescence improves grain protein, zinc, and iron content in wheat, Science, 2006, 314, 1298–1301 CrossRef CAS PubMed.
- H. S. Randhawa, et al., Application of molecular markers to wheat breeding in Canada, Plant Breed., 2013, 132, 458–471 CAS.
- M. W. Blair and P. Izquierdo, Use of the advanced backcross-QTL method to transfer seed mineral accumulation nutrition traits from wild to Andean cultivated common beans, Theor. Appl. Genet., 2012, 125, 1015–1031 CrossRef PubMed.
- B. S. Kodkany, et al., Biofortification of pearl millet with iron and zinc in a randomized controlled trial increases absorption of these minerals above physiologic requirements in young children, J. Nutr., 2013, 143, 1489–1493 CrossRef CAS PubMed.
- J. L. Finkelstein, et al., A randomized trial of iron-biofortified pearl millet in school children in India, J. Nutr., 2015, 145, 1576–1581 CrossRef CAS PubMed.
- J. D. Haas, et al., Consuming iron biofortified beans increases iron status in Rwandan women after 128 days in a randomized controlled feeding trial, J. Nutr., 2016, 146, 1586–1592 CrossRef CAS PubMed.
- A. A. T. Johnson, et al., Constitutive overexpression of the OsNAS gene family reveals single-gene strategies for effective iron- and zinc-biofortification of rice endosperm, PLoS One, 2011, 6, e24476 CAS.
- N. Narayanan, et al., Overexpression of Arabidopsis VIT1 increases accumulation of iron in cassava roots and stems, Plant Sci., 2015, 240, 170–181 CrossRef CAS PubMed.
- S. Borg, et al., Wheat ferritins: Improving the iron content of the wheat grain, J. Cereal Sci., 2012, 56, 204–213 CrossRef CAS.
- J. M. Connorton, et al., Functional analysis of vacuolar iron transporters in wheat (Triticum aestivum) identifies TaVIT2 as an iron and manganese transporter effective for biofortification of cereals, Plant Physiol., 2017 Search PubMed , accepted.
- I. H. Slamet-Loedin, S. E. Johnson-Beebout, S. Impa and N. Tsakirpaloglou, Enriching rice with Zn and Fe while minimizing Cd risk, Front. Plant Sci., 2015, 6, 1–9 Search PubMed.
- R. Banakar, A. Álvarez Fernández, J. Abadía, T. Capell and P. Christou, The expression of heterologous Fe (III) phytosiderophore transporter HvYS1 in rice increases Fe uptake, translocation and seed loading and excludes heavy metals by selective Fe transport, Plant Biotechnol. J., 2016, 15, 423–432 CrossRef PubMed.
- K. V. Krasileva, et al., Uncovering hidden variation in polyploid wheat, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, E913–E921 CrossRef CAS PubMed.
- R. Hurrell and I. Egli, Iron bioavailability and dietary reference values, Am. J. Clin. Nutr., 2010, 91, 1461S–1467S CrossRef CAS PubMed.
- V. Raboy, K. A. Young, J. A. Dorsch and A. Cook, Genetics and breeding of seed phosphorus and phytic acid, J. Plant Physiol., 2001, 158, 489–497 CrossRef CAS.
- J. Wirth, et al., Rice endosperm iron biofortification by targeted and synergistic action of nicotianamine synthase and ferritin, Plant Biotechnol. J., 2009, 7, 631–644 CrossRef CAS PubMed.
- N. Yamaji, et al., Reducing phosphorus accumulation in rice grains with an impaired transporter in the node, Nature, 2017, 541, 92–95 CrossRef CAS PubMed.
- T. Eagling, A. A. Wawer, P. R. Shewry, F. Zhao and S. J. Fairweather-tait, Iron bioavailability in two commercial cultivars of wheat: comparison between wholegrain and white flour and the effects of nicotianamine and 2′-deoxymugineic acid on iron uptake into Caco-2 cells, J. Agric. Food Chem., 2014, 62, 10320–10325 CrossRef CAS PubMed.
- F. Leenhardt, M. A. Levrat-Verny, E. Chanliaud and C. Rémésy, Moderate decrease of pH by sourdough fermentation is sufficient to reduce phytate content of whole wheat flour through endogenous phytase activity, J. Agric. Food Chem., 2005, 53, 98–102 CrossRef CAS PubMed.
- G. O. Latunde-Dada, et al., Micromilling enhances iron bioaccessibility from wholegrain wheat, J. Agric. Food Chem., 2014, 62, 11222–11227 CrossRef CAS PubMed.
- Y. Kakei, et al., Development of a novel prediction method of cis-elements to hypothesize collaborative functions of cis-element pairs in iron-deficient rice, Rice, 2013, 6, 22 CrossRef PubMed.
- A. Koryachko, et al., Clustering and Differential Alignment Algorithm: Identification of early stage regulators in the Arabidopsis thaliana iron deficiency response, PLoS One, 2015, 10, 1–21 Search PubMed.
- H.-J. Mai, S. Pateyron and P. Bauer, Iron homeostasis in Arabidopsis thaliana: transcriptomic analyses reveal novel FIT-regulated genes, iron deficiency marker genes and functional gene networks, BMC Plant Biol., 2016, 16, 211 CrossRef PubMed.
- T. Kobayashi, et al., The rice transcription factor IDEF1 is essential for the early response to iron deficiency, and induces vegetative expression of late embryogenesis abundant genes, Plant J., 2009, 60, 948–961 CrossRef CAS PubMed.
- Y. Ogo, et al., A novel NAC transcription factor, IDEF2, that recognizes the Iron Deficiency-Responsive Element 2 regulates the genes involved in iron homeostasis in plants, J. Biol. Chem., 2008, 283, 13407–13417 CrossRef CAS PubMed.
- K. R. Trijatmiko, et al., Biofortified indica rice attains iron and zinc nutrition dietary targets in the field, Sci. Rep., 2016, 6, 19792 CrossRef CAS PubMed.
- S. P. Singh, B. Keller, W. Gruissem and N. K. Bhullar, Rice NICOTIANAMINE SYNTHASE 2 expression improves dietary iron and zinc levels in wheat, Theor. Appl. Genet., 2017, 130, 283–292 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2017 |