On-resin synthesis of Lanreotide epimers and studies of their structure–activity relationships†
Arnab
Chowdhury
 a,
Nitesh Mani
Tripathi
a,
Nitesh Mani
Tripathi
 a,
Rohit
Jadav
b,
Vinod
Gour
a,
Parva
Purohit
b and
Anupam
Bandyopadhyay
a,
Rohit
Jadav
b,
Vinod
Gour
a,
Parva
Purohit
b and
Anupam
Bandyopadhyay
 *a
*a
aBiomimetic Peptide Engineering Laboratory, Department of Chemistry, Indian Institute of Technology Ropar, Rupnagar, Punjab-140001, India. E-mail: anupamba@iitrpr.ac.in
bKashiv BioSciences Pvt Ltd., Block-B, Sardar Patel Ring Rd, opp. Applewoods Township, Ahmedabad, Gujarat-382210, India
First published on 1st July 2024
Abstract
Peptide drugs often accompany epimeric impurities (isomers). Therefore, efficient chemical synthesis of epimers is critical to identify them correctly and investigate their biological activities. Here, we report the rapid synthesis and structure–activity relationship (SAR) studies of eight possible epimers of a somatostatin synthetic analog (SSA), lanreotide (LAN). SPPS and the subsequent on-resin rapid disulfide closure method offered >90% conversion yield for all epimers (P1–P8). Further, we developed an analytical method to separate these epimers, which enabled the profiling of five epimeric impurities in the API, purchased for Somatuline generic formulations. In SAR studies, most LAN epimers revealed compromised antiproliferative activity, while the P7 epimer retained antiproliferative activity similar to LAN API, as supported by in silico SAR studies in detail. Additionally, P7 showed serum stability nearly identical to LAN, suggesting that drug epimers could be a potential API. Current studies will further encourage the development of novel SSA scaffolds.
Introduction
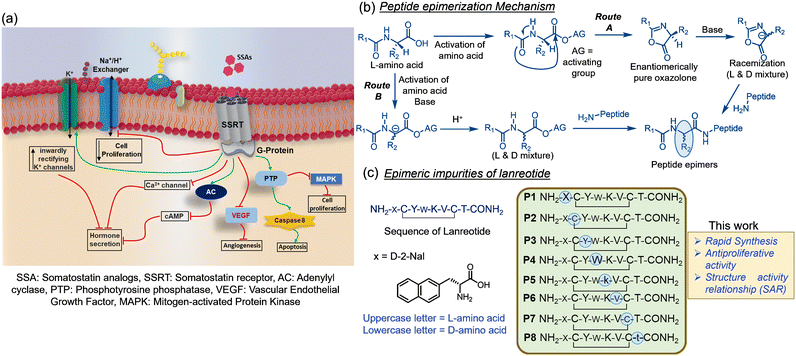
Somatostatin analogs (SSA) are a widely used therapeutic option in treating somatotropinomas, thyrotropinomas, and gastroenteropancreatic neuroendocrine tumors.1 SSA are known to encompass direct and indirect mechanisms for antiproliferative activity. Cell cycle arrest and/or modulation of apoptosis are the direct mechanisms of action. In contrast, inhibition of several growth and proangiogenic factors' secretions is indirectly associated with activities upon binding to somatostatin receptors (SSRT).2,3 The primary antiproliferative signaling pathways are represented in Fig. 1a. Among clinically used SSA, lanreotide (LAN) is a potent synthetic analog of human somatostatin, marketed under the brand name Somatuline for the management of acromegaly4,5 and the treatment of neuroendocrine tumors.5 The current census of cancer progression demands the generic production of SSA for societal wellness. However, a generic formulation of peptide drugs mandates rigorous impurity profiling, safety evaluation, and adherence to sameness with marketed reference-listed products (RLD) to avoid side effects.6–9Peptide drug impurities frequently originate during the synthesis due to side reactions,10 residue truncation, and racemization of activated amino acids (Fig. 1b) and formulation.11 Identification of drug-related impurities is highly rely on their efficient synthesis and analytical validation. Most SSAs contain a disulfide bond in the peptide chain. Therefore, synthesizing their related impurities – extends a step of disulfide assembly in the solution phase by the popular methods, air, iodine, and DMSO oxidation.12,13 In contrast, disulfide assembly in the solid phase is more advantageous.13–15 We have recently developed highly efficient and rapid iodine-mediated oxidative disulfide assembly using a persulfate (S2O82−) additive15 on resin to eliminate the laborious process in the solution phase.
Here, we sought to exploit I2-S2O82− mediated on resin disulfide assembly for synthesizing all eight epimers of LAN (Fig. 1c) to know the method's suitability in producing a reasonable quantity (∼0.5 g) that can be utilized in impurity profiling and biological activity studies for the generic formulation of Somatuline in the industry. Towards this end, the method delivered excellent disulfide assembly transformation for all possible epimers (P1–P8), which enabled hassle-free purifications and isolations of these epimers and profiling epimer impurities coexist with purchased LAN API. We further extended our interest in understanding epimers' in vitro and in silico structure–activity relationships, which may render the scope for developing novel peptide scaffolds. In due course, we found that P7 exhibited a similar activity and serum stability profile to LAN API. Structure–activity relationship studies in silico revealed that P7 enjoys interaction modules akin to LAN by forming a thermodynamically stable complex with SSR2 receptors.
Results and discussion
Epimeric impurities are a common occurrence in recent practices of peptide synthesis. During the synthesis, the activation of each amino acid always produces a certain amount of racemized oxazolone or racemized activated ester, which leads to epimeric side products upon amidation with the peptide chain (Fig. 1b). Most epimeric impurities pose a challenging task to separate on a preparative scale due to subtle changes in their chemical properties compared to the API (active pharmaceutical ingredients) peptide. Therefore, it is crucial to synthesize all epimeric isomers to profile the impurities coexisting with an API before generic formulations. Theoretically, eight epimers can be generated in the chemical synthesis of LAN (Fig. 1c) if the racemization of a single amino acid at a time in the peptide sequence is concerned. Isomers bearing multiple racemized residues are believed to be way below the regulatory limit of impurity (<0.1%), and thus, they are not under consideration.All linear peptide chains were elongated through conventional solid-phase peptide synthesis (SPPS) with Fmoc chemistry on rink-amide polystyrene resin (0.8 mmol). HBTU was used as a coupling agent (ESI† section II) in the SPPS. Then, the resin was treated with I2-S2O82− in DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water (9
water (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
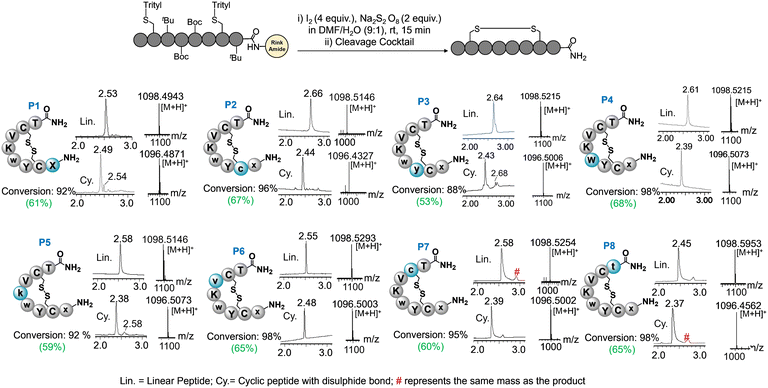
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) systems for disulfide assembly.15 The reaction offered excellent disulfide conversion for all epimers in 15 min, confirmed by the LC-MS analysis (Fig. 2, ESI† section III and Fig. S1–S8). In contrast, the solution phase disulfide assembly with I2 often requires highly dilute conditions and hours (>3–5 h) to complete the reaction. This reaction also requires the quenching and elimination of I2 at the end to avoid side products on a large scale,16 which makes the process laborious and time-consuming. Solid-phase disulfide assembly is highly advantageous for economical and timely production. The method with I2-S2O82− showed no side products in solid-phase disulfide assembly. In the case of crude P8, a slight hump was observed near the major peak also contained the product mass, which needs further investigation. Epimers were cleaved from resin, purified through HPLC, and lyophilized to receive white powder (∼0.5 g per each) ESI† section IV.
1) systems for disulfide assembly.15 The reaction offered excellent disulfide conversion for all epimers in 15 min, confirmed by the LC-MS analysis (Fig. 2, ESI† section III and Fig. S1–S8). In contrast, the solution phase disulfide assembly with I2 often requires highly dilute conditions and hours (>3–5 h) to complete the reaction. This reaction also requires the quenching and elimination of I2 at the end to avoid side products on a large scale,16 which makes the process laborious and time-consuming. Solid-phase disulfide assembly is highly advantageous for economical and timely production. The method with I2-S2O82− showed no side products in solid-phase disulfide assembly. In the case of crude P8, a slight hump was observed near the major peak also contained the product mass, which needs further investigation. Epimers were cleaved from resin, purified through HPLC, and lyophilized to receive white powder (∼0.5 g per each) ESI† section IV.
Next, we inspected the identity of epimeric impurities accompanied by the purchased LAN API. An HPLC method was developed with a gradient system using the water/acetonitrile (0.1% formic acid) buffer for two hours to separate eight epimer impurities, including LAN API. Except for P7, the developed HPLC1 method enabled distinct epimers' separation with LAN API in the total run time (ESI† section Ie, Fig. S9), which further facilitated the identification of epimeric impurities and estimated their amount (%) in the purchased API. P7 eluted immediately after the LAN-API, which led to trouble quantifying this epimer. A distinct method, HPLC2, was also developed in this setting to obtain correct quantitative estimations of P7 (ESI† section Ie, Fig. S9g). We found twelve coexisting impurities in the purchased API separated in the HPLC1 method. Among them, the retention time of five epimers (P1, P3–P5, P7) matched (Fig. 3a). Noteworthy, each epimer measured ∼0.3% occupancy in the purchase API, which is above the regulatory limit of an impurity that needs to be considered complete biological profile studies before passing to generic formulations. The rest of the epimeric impurities did not appear in the impurity profile; perhaps they coexisted below the detection limit or were separated out during the HPLC purification of API. Moreover, the impurity profile demonstrated the coexistence of the five potential epimeric impurities above the regulatory amount with the batch of purchased LAN API.
Further, we were curious to explore the antiproliferative activity of epimers and understand their activity modulation with the change of a chiral center. In order to test, eight epimers and LAN API were incubated with A549 at 20 μM for 12 hours. A549 is a lung carcinoma epithelial cell line reported to highly express SSR2 receptors.17 After incubation, an MTT assay was performed to assess cell viability and compare outcomes with LAN API (Fig. 3b, ESI† section V). LAN is a well-preorganized peptide scaffold; thus, altering a single amino acid's absolute configuration in the sequence is expected to compromise the original activity. As anticipated, most epimers illustrated activity compromisation in the MTT assay. However, P4 and P7 epimers retained apoptotic activity similar to LAN.
Interestingly, no epimer showed a complete cessation of apoptotic activity. P2 showed the highest cell viability among all epimers, demonstrating the maximum activity compromise with altering the chiral center at 2nd residue. Throughout the biological experiments, untreated and cisplatin-treated cells were considered as controls. MTT results encouraged us to further study a dose-dependent cell viability assay with P2, P4, P7, and LAN API for measuring half-maximal inhibitory concentration (IC50). A series of concentrations, starting with 5 nM to 1000 nM, were incubated with A549 cells for 48 hours (Fig. 3b, ESI† section V and VI). The prolonged 48 hours cell viability assay proved that the IC50 value of P7 (∼25 nM) is close to LAN API (∼21 nM), while the IC50 value of P4 (∼40 nM) is slightly higher than that of LAN. Nonetheless, epimerization at Cys2 residue (P2) negatively impacts apoptotic activity, further manifested by the IC50 value, which is around six times higher than LAN API.
The experimented apoptotic activity of P4 and P7 epimers further motivated us to test their serum and trypsin stability and compare them with LAN. A stock solution of P4, P7, and LAN was individually mixed with 25% human fresh blood serum diluted in PBS (1×) at ∼250 μM final peptide concentration and incubated for 24 hours at 37 °C. The linear precursor of LAN peptide was included as a positive control because it is speculated to degrade faster than the cyclic version. Then, the degradation of each peptide in serum was monitored via HPLC at specific intervals. As projected, the linear LAN precursor showed 50% degradation within 15 hours. P4 was 40% degraded after 24 hours (Fig. 3c, ESI† section VII), however, LAN and P7 showed only 4% and 10% degradation, respectively. Although P4 showed great antiproliferative activity, its serum stability is poor because of L-Trp at the 4th residue. When these four peptides were subjected to trypsin digestion, degradation profiles revealed parallel stability profiles to human blood serum (Fig. 3d, ESI† section VIII). These data collectively represent a comparable antiproliferative activity and stability profile of P7 and LAN peptides. We anticipated that P7 would display a better stability profile than LAN because of D-cys at the 7th residue. Perhaps P7 acquires relatively more conformational strain than LAN, which results in a relatively higher population of its linear precursor (reduced disulfide) at physiological conditions,18 thus realizing a slightly more proteolytic cleavage.
The cell antiproliferative activities shown by P4, P7, and LAN were further quantitatively assessed for a final judgment. Cell apoptosis events were monitored in a flow cytometer by labeling externalized phosphatidylserine with annexin V-FITC (ESI† section IX, Fig. S18). Briefly, A549 cells were incubated with 20 μM peptide for 12 hours and treated with annexin V-FITC for flow cytometry assessment. The fluorescence readouts in the flow cytometer (Fig. 3e) stood consistent with the outcome demonstrated in the MTT assay. The bar graph and histogram data are triplicated median values of FITC-labeled apoptotic cell populations (see ESI† Fig. S17 for cell population quadrants). Put together, altering the chiral center at 4th (4w/W) and 7th (7C/c) residue abided comparable activity to LAN API.
By looking at the apoptotic activity profiles of epimers, we were curious to understand why P4 and P7 epimers show similar activity to LAN. We conducted an in silico docking and molecular dynamics (MD) simulation to reveal SAR. Since LAN shows high selectivity in binding with SSR2, the crystal structure of SSR2-LAN (PDB 7XAV)19 complex was used for receptor/protein preparation (Fig. 4a, ESI† section X). Targeted docking was performed at the binding site of LAN using Glide/XP after preparing the epimer ligands P2, P4, and P7 with the LigPrep module in Schrodinger.20,21 We included P2 in these SAR studies to reveal the insight of binding compromise, and it would also serve as another control along with LAN. The best poses with the lowest docking score were further subjected to an MD Simulation (100 ns) using Desmond22 to assess the thermodynamic stability of the complexes (ESI section Xd). RMSD, RMSF, and per residue interaction analyses were performed with the post-MD simulation of docked complexes, which authenticated further thermodynamic stability of the chosen docked complexes throughout the simulated trajectories.
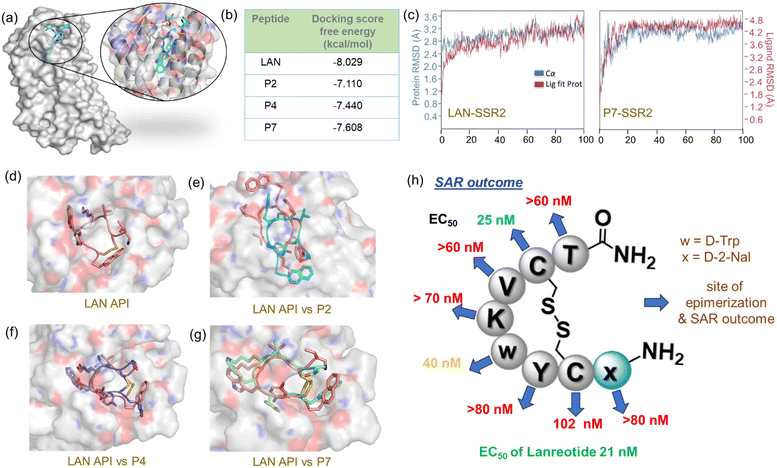 | ||
| Fig. 4 (a) Crystal structure of SSR2 bound to LAN. The binding site is zoomed in for better inspection of the binding site. (b) Glide docking binding free energy score between SSR2 and each epimer. The score for LAN served as a control, (c) comparison of RMSD calculation between LAN-SSR2 and P7-SSR2 complex. (d) The docked structure between SSR2 and LAN. A comparison study of SSR2 binding posture by superimposition between LAN and (e) P2, (f) P4, and (g) P7. Py-MOL,23 as a molecular graphics system, was used for visualization. (h) A summarized picture of Lanreotide epimers' SAR. Blue arrows represent residue epimerization and activity outcome of epimers. Green (slight), orange (medium), and red (compromised) EC50 outcomes represent the deviation of activity concerning LAN API. | ||
The in silico ligand binding free energy pattern (Fig. 4b) demonstrated consistent results with the experimental outcome, demonstrating that SSR2-LAN provides the most stable complex than considered epimers. The analyses of interaction modules in simulated SSR2-LAN complex showed a maximum ligand interaction occupancy with key SSR2 residues, Asp122, Gln126, Trp197, Asn276, Ser282, Asp295, and Tyr302 (ESI† section Xe, Fig. S23). During the MD simulation, we observed that a bridging water molecule and hydrogen bonds remained intact in the SSR2-LAN complex, signifying the formation of a high-affinity complex. Analogous interaction patterns were observed in the case of the SSR2-P7 complex, but a few interactions with P4 were perturbed due to the change in absolute configuration at the Trp residue (Fig. S25–S26†). Contrarily, the SSR2-P2 complex depicts an entirely different interaction with a 180° rotated binding posture, perhaps the reason for compromising antiproliferative activity (Fig. 4e).
We did not observe any significant variation in the RMSD values of LAN, P4, and P7 complex with SSR2 receptor, which integrated stable conformation and complex in their docked position (Fig. 4c, ESI† Fig. S19). However, abnormal changes in P2-SSR2 RMSD values were observed as expected, which can be correlated with the experimented maximum cell viability. All interaction details are elaborated on and shown in the ESI† section Xe. Each epimer (P2, P4, and P7) binding posture to the SSR2 receptor is superimposed with the LAN-SSR2 complex for better comparison (Fig. 4d–f). In vitro and in silico SAR studies illustrated (Fig. 4h) that LAN is a highly optimized ligand for selective and tight binding to SSR2 receptor. Because perturbation of its 3D-preorganized structure by any means compromises the SSR2 binding and SSTR signaling process. In this study, the activity and stability profile of P7 intrigued us to investigate drug epimers that are equally potential for drug development. We believe these studies may help understand the SAR of peptide drug epimers and explore novel molecules that are selective for other SSRs.
Conclusion
We have established a complete on-resin synthesis route for all LAN epimers, which included a highly efficient step of I2-S2O82−-mediated disulfide assembly. Further, synthetic epimers helped identify and validate five epimeric impurities coexisting with purchased LAN API through newly developed HPLC methods. SAR studies revealed that most epimers compromised the original activity of LAN but abided by a fair extent of antiproliferative activity. The epimer P7, which has D-cys at the 7th residue, displayed a subtle change in original antiproliferative activity and high proteolytic stability in serum akin to LAN. The in silico SAR studies have further confirmed that P7 attributes similar binding interactions to SSR2 as LAN, which is the primary reason for retaining such apoptotic activity. SAR studies concluded that LAN epimers could show similar apoptotic activity if they adopt a nearly identical 3D preorganized structure of API. In this regard, changes in the hot-spot residue configuration are shown to compromise activity but not abrogate binding. Altering the configuration at the Cys2 residue resulted in the worst binding epimer because of the high deviation to the 3D architecture of API. This work advocates that drug epimers could be potential candidates for drug development against different receptor subtypes and biological targets.Notes
All experiments were performed in accordance with the guidelines of DBT-Biosafety, and experiments were approved by the Institutional Biosafety Committee (IBSC) at IIT Ropar. Informed consents were obtained from human participants of this study.Data availability statements
The data supporting this article have been included as part of the ESI.†Author contributions
AB conceived the project. VG and AC synthesized, isolated, and analyzed all compounds with the guidance of AB. NMT performed all in silico studies. RJ and PP performed HPLC analysis and epimer impurity profiling. The manuscript was written with contributions from all authors. All authors have approved the final version of the manuscript.Conflicts of interest
AB and VG have filed an Indian patent application (IN202211063074) for on-resin disulfide assembly.Acknowledgements
We acknowledge IIT Ropar for the financial support and Kashiv Bioscience Pvt. Ltd., India, for the Lanreotide project. We thank CRF IIT Ropar for the NMR and HRMS facility and FIST-DST (SR/FST/CS-I/2018/55) for departmental mass. The authors thank Schrödinger, LLC for providing the trial license to perform the computational study to carry out this work. All computations were carried out using the IIT Ropar HPC facility.References
- M. Chalabi, C. Duluc, P. Caron, D. Vezzosi, J. Guillermet-Guibert, S. Pyronnet and C. Bousquet, Trends Endocrinol. Metab., 2014, 25, 115–127 CrossRef CAS
.
- M. C. Cantone, A. Dicitore and G. Vitale, J. Clin. Med., 2021, 10, 501 CrossRef CAS PubMed
.
- F. Barbieri, A. Bajetto, A. Pattarozzi, M. Gatti, R. Würth, S. Thellung, A. Corsaro, V. Villa, M. Nizzari and T. Florio, Int. J. Pept., 2013, 2013, 20 Search PubMed
.
- Somatuline Autogel 60 mg, solution for injection in a prefilled syringe - Summary of Product Characteristics (SmPC) - (emc), https://www.medicines.org.uk/emc/product/4808/smpc#gref, (accessed 2 May 2023).
- E. M. Wolin, A. Manon, C. Chassaing, A. Lewis, L. Bertocchi, J. Richard and A. T. Phan, J. Gastrointest. Cancer, 2016, 47, 366–374 CrossRef CAS PubMed
.
- L. Wu, RSC Drug Discovery Ser., 2019, 2019, 1–30 Search PubMed
.
- D. Zane, P. L. Feldman, T. Sawyer, Z. Sobol and J. Hawes, Internet J. Toxicol., 2021, 40, 108–124 CrossRef CAS
.
- V. Vergote, C. Burvenich, C. Van De Wiele and B. De Spiegeleer, J. Pept. Sci., 2009, 15, 697–710 CrossRef CAS PubMed
.
- J. B. Biomed, S. Van Dorpe, M. Verbeken, E. Wynendaele and B. De Spiegeleer, J. Bioanal. Biomed., 2011, S6, 003 Search PubMed
.
- S. Chatterjee and A. Bandyopadhyay, J. Pept. Sci., 2023, 29, e3489 CrossRef CAS PubMed
.
- R. Jadav, R. Kameriya, S. Chatterjee, V. Gour, P. Purohit and A. Bandyopadhyay, J. Pept. Sci., 2024, 30, e3564 CrossRef CAS PubMed
.
- R. He, J. Pan, J. P. Mayer and F. Liu, ChemBioChem, 2020, 21, 1101–1111 CrossRef CAS
.
- K. Kobayashi, A. Taguchi, Y. Cui, H. Shida, K. Muguruma, K. Takayama, A. Taniguchi and Y. Hayashi, Eur. J. Org. Chem., 2021, 2021, 956–963 CrossRef CAS
.
- T. M. Postma and F. Albericio, Org. Lett., 2013, 15, 616–619 CrossRef CAS
.
- A. Chowdhury, V. Gour, B. K. Das, S. Chatterjee and A. Bandyopadhyay, Org. Lett., 2023, 25, 1280–1284 CrossRef CAS
.
- K. M. B. Reddy, Y. B. Kumari, D. Mallikharjunasarma, K. Bulliraju, V. Sreelatha and K. Ananda, Int. J. Pept., 2012, 2012, 323907 Search PubMed
.
- J. Fan, L. Deng, Y. Peng and Y. Ding, Sci. Rep., 2022, 12, 1–12 CrossRef
.
- R. Zhang and G. H. Snyder, J. Biol. Chem., 1989, 264, 18472–18479 CrossRef CAS
.
- Q. Bo, F. Yang, Y. Li, X. Meng, H. Zhang, Y. Zhou, S. Ling, D. Sun, P. Lv, L. Liu and C. Tian, Cell Discovery, 2022, 8, 47 CrossRef CAS
.
- Schrödinger Release 2021: LigPrep, Schrödinger, LLC, New York, NY, 2021 Search PubMed.
- Maestro-Desmond Interoperability Tools, Schrödinger, New York, NY, 2021 Search PubMed.
- Desmond Molecular Dynamics System, D. E. Shaw Research, New York, NY, 2021 Search PubMed.
- The PyMOL Molecular Graphics System, Version 2.5, Schrödinger, LLC Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4md00338a |
| This journal is © The Royal Society of Chemistry 2024 |