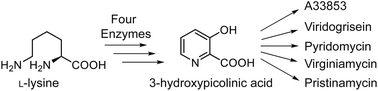
3-Hydroxypicolinic acid (3-HPA) is an important pyridine building block of bacterial secondary metabolites. Although the main biosynthetic pathways of these metabolites have been identified and well characterized, the enzymatic mechanism underlying the biosynthesis of 3-HPA has yet to be elucidated. In this work, we successfully reconstituted the complete biosynthetic pathway of 3-HPA in vitro. We showed that an L-lysine 2-aminotransferase, a two-component monooxygenase, and a FAD-dependent dehydrogenase are required to convert L-lysine to 3-HPA. We further demonstrated that 3-HPA does not derive from the direct hydroxylation of the picolinic acid at C-3, but from a successive process of C-3 hydroxylation of the piperideine-2-carboxylic acid and tautomerization of the produced 3-hydroxyl dihydropicolinic acid. Therefore, this study unveils the unusual assembly logic of 3-HPA and sheds light on the potential of engineering the 3-HPA pathway for generating novel pyridine-based building blocks.