 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Gold nanoparticle-assisted polymerase chain reaction: effects of surface ligands, nanoparticle shape and material†
Ekaterina
Vanzha
a,
Timofey
Pylaev
a,
Vitaly
Khanadeev
a,
Svetlana
Konnova
b,
Valentina
Fedorova
b and
Nikolai
Khlebtsov
 *ac
*ac
aInstitute of Biochemistry and Physiology of Plants and Microorganisms, Russian Academy of Sciences, 13 Prospekt Entuziastov, Saratov 410049, Russia. E-mail: khlebtsov@ibppm.ru
bSaratov Scientific and Research Veterinary Institute, Russian Academy of Agricultural Sciences, 6 Ulitsa 53 Strelkovoj Divizii, Saratov 410028, Russia
cSaratov National Research State University, 83 Ulitsa Astrakhanskaya, Saratov 410012, Russia
First published on 14th November 2016
Abstract
Recent reports have shown significant improvement of the polymerase chain reaction (PCR) efficiency and specificity with the addition of gold nanoparticles (AuNPs). However, the underlying mechanisms are still unclear and debatable. Here, we report effects of AuNP concentration, shape, material, and surface functionalization on PCR outcome under typical amplification conditions. All nanoparticles were tested with two PCR diagnostic models: (1) the nitrogen fixation (NifD) gene from Azospirillum brasilense Sp7 bacteria and (2) the polymorphic ompA gene encoding major outer membrane protein of Chlamydia trachomatis. The optimal concentration of 16 nm citrate-stabilized negatively charged spherical AuNPs (zeta potential −29.9 mV) was determined to be 0.4 nM, in agreement with previous data for quite different PCR systems. Functionalization with poly(vinylpyrrolidone) (PVP) and thiolated PEG (mPEG-SH) essentially did not change the zeta-potential (−30 mV and −21 mV, respectively), but completely suppressed the PCR enhancement. Positively charged AuNPs with poly(diallyldimethylammonium) (PDDA) and cetyltrimethylammonium bromide (CTAB) capping polymers exhibited no specific enhancing effect on PCR or even inhibited the reaction at 5 pM of AuNP–PDDA. Also, no specific PCR enhancement was obtained with CTAB-stabilized gold nanorods. To elucidate the role of the particle material, we performed PCR with gold and silica particles of comparable size and equal negative charges. In contrast to the great PCR enhancement with 16 nm AuNPs, 20 nm silica nanoparticles did not affect the PCR efficiency or inhibit the reaction at concentrations higher than 100 nM. These findings suggest that the nature of the PCR system components, surface stabilizing agents and the thermal conductivity of the nanoparticle material can play important roles in the nanoparticle-mediated PCR.
1. Introduction
The polymerase chain reaction (PCR) has become a standard biomolecular technique, which is commonly used for sensitive and rapid DNA detection. Exponential amplification of PCR allows replication of a single target copy to produce a large amount of DNA copies.1–3 However, the PCR can be error-prone and the specificity and efficiency of PCR can be unsatisfactory without proper optimization.4 As a result of the poor purification of template DNA, primer–template mismatches, spontaneous formation of primer dimers, and non-optimized PCR-mixture ratio and thermocycling conditions, the formation of the target amplicon may be accompanied by non-specific side products called PCR artifacts. To fix this problem, several enhancers have been introduced: (1) various additives in PCR mixture, such as betaine,5 dithiothreitol,6 dimethyl sulfoxide,7 single-stranded DNA-binding protein (SSB);8 (2) instrumental design, including development of thermocyclers with precise heating/cooling rates;9,10 (3) optimization of PCR system through optimal Mg2+ concentration, cycle numbers and proper primer design;11–13 (4) enzyme modification;14,15 and (5) new touchdown16 and nested17 PCR strategies. Although these tools improve the PCR outcome, yet they are not all-purpose and the optimization protocol can be case dependent.During the past decade, various nanomaterials have been used for PCR enhancement, including gold nanoparticles (AuNPs),18 silver nanoparticles and graphene oxide,19 reduced graphene oxide,19 titanium dioxide and quantum dots,20 upconversion nanoparticles,21 fullerenes (C60),22 carbon nanoparticles and nanotubes,23 carbon nanopowder,24 magnetic nanoparticles,25 semiconductor nanomaterials,26,27 dendrimers,28etc. Nevertheless, AuNPs are one of the promising candidates for use as PCR enhancers owing to their controlled geometrical parameters, reproducible synthesis and functionalization protocols. Recently, several hypothetic mechanisms of AuNP-assisted PCR were proposed to account for: (1) the SSB-like mechanism29 and electrostatic interactions between AuNPs and PCR components;30 (2) the high surface-to-volume ratio and non-specific adsorption of PCR components;28–31 and (3) the thermal properties of AuNPs.32 However, none of the suggested mechanisms can be considered convincing.
Here, we elucidated the role of the particle functionalization with neutral (PVP and mPEG-SH) and cationic (PDDA and CTAB) polymer ligands and role of the nanoparticle material by performing PCR with gold and silica particles of comparable size and charge, other PCR conditions being unchanged. In addition, the nanoparticle shape effect was examined with CTAB stabilized nanorods. All nanoparticles were tested with two PCR diagnostic models: (1) the nitrogen fixation (NifD) gene from Azospirillum brasilense Sp7 bacteria; and (2) the polymorphic ompA gene encoding major outer membrane protein of Chlamydia trachomatis. At an optimal concentration of citrate-stabilized 16 nm negatively charged spherical AuNPs (0.4 nM), the PCR specificity and efficiency was greatly enhanced, whereas the negatively charged PVP- and PEG-stabilized AuNPs did not reveal any enhancing properties. Also, no specific PCR enhancement was observed with positively charged AuNPs and gold nanorods. By contrast to 16 nm negatively charged citrate-stabilized AuNPs, silica nanoparticles with the same charge and comparable 20 nm size have no impact on PCR efficiency. These observations show an important role of the nanoparticle material and surface modification in PCR enhancement, and could be helpful for further mechanistic studies.
2. Results and discussion
2.1 Characterization of nanoparticles
The following as-prepared and surface-functionalized AuNPs were used: negatively charged citrate-stabilized spherical gold nanoparticles (AuNP); nanoparticles capped with neutral polymers PVP and mPEG-SH (AuNP–PVP and AuNP–PEG-SH, respectively) and cationic polymers CTAB and PDDA (AuNP–CTAB and AuNP–PDDA, respectively); gold nanorods (AuNR-662, 662 stands for the longitudinal plasmon resonance maximum wavelength) with aspect ratio of about 2.3; and spherical silica nanoparticles (SiO2NP). The acronym definitions and chemical structures of polymeric ligands are given in ESI file (Section S1).†Fig. 1 shows illustrative TEM images of citrate-stabilized AuNPs, gold nanorods, and silica nanospheres. Extinction spectra of nanoparticles are shown in Fig. S1 (ESI).†Table 1 summarizes the values of the zeta-potentials ς (mV), absorption maximum wavelengths of as-prepared NPs in aqueous solution (for nanorods, the longitudinal resonance is indicated); the average TEM and DLS sizes; the nanoparticle molar concentrations normalized to the extinction Amax = 1; and the optimal concentrations and maximal specificity and efficiency of different PCR additives used in our PCR systems. | ||
| Fig. 1 Representative TEM images of as-prepared AuNPs (a), gold nanorods (b), and silica nanoparticles (c). | ||
| Sample | Zeta potential (mV) | TEM size (nm) | DLS size (nm) | λ max (nm) | Concentration (nM) | Optimal concentration | Maximal efficiency | Maximal specificity |
|---|---|---|---|---|---|---|---|---|
| AuNP | −29.9 ± 3 | 16 ± 0.8 | 17.1 ± 1.2 | 520 | 4 | 0.4 nM | 4.25 ± 0.33 | 5.7 ± 0.43 |
| AuNP–PVP | −30 ± 3 | 16 ± 1.0 | 22.8 ± 1.8 | 522 | 4 | 0.4 nM | 0.82 ± 0.07 | 1.07 ± 0.11 |
| AuNP–PEG-SH | −20.5 ± 3 | 16 ± 1.0 | 26.4 ± 1.3 | 522 | 4 | 4 nM | 0.95 ± 0.08 | 1.09 ± 0.12 |
| AuNP–CTAB | +10.2 ± 1 | 25 ± 1.2 | 28.6 ± 2.3 | 524 | 4 | 1 fM | 1.69 ± 0.14 | 1.03 ± 0.14 |
| AuNP–PDDA | +30.5 ± 2 | 16 ± 1.0 | 23.4 ± 1.7 | 522 | 2.8 | 1 fM | 2.69 ± 0.13 | 1.5 ± 0.11 |
| AuNR-662 | +44.7 ± 2 | d = 25.1 ± 1.5, L = 70.2 ± 7.8 | 63.3 ± 6.8 | 657 | 0.26 | 1 fM | 0.9 ± 0.07 | 1.07 ± 0.13 |
| SiO2NP | −31.2 ± 3 | — | 20.0 ± 2.5 | — | 490 | 1 nM | 0.97 ± 0.08 | 1.07 ± 0.12 |
It follows from Table 1 that the average TEM and DLS diameters of citrate NPs (16 ± 0.8 nm and 17 ± 1.2 nm, respectively) have no significant difference within the standard deviation (SD) of about 1 nm. By contrast, after functionalization with PVP, PEG-SH, CTAB, and PDDA polymeric ligands, the hydrodynamic diameters increase significantly by 6.8, 10.4, 3.6, and 7.4 nm. We consider these data as strong evidence for successful attachment of ligands. The second evidence comes from drastic change in colloidal stability of NPs before and after functionalization. Fig. S2 (ESI†) shows extinction spectra and photos of citrate-stabilized and ligand-stabilized AuNPs before and after addition of 0.1 M NaCl salt. As expected, the addition of salt to the citrate-stabilized AuNPs results in immediate aggregation which is evident from changes in suspension color and extinction spectrum. By contrast, the addition of the same amount of salt to the ligand-stabilized AuNPs does not induce any aggregation phenomena accompanied by color and spectral changes. What is more, the ligand-stabilized NPs do not aggregate even in salt environment under strong temperature variations during PCR cycles (see below, Fig. 4).
2.2 Mechanisms of AuNP enhancing effect
Effect of nanoparticles on non-optimized PCR system was investigated by performing PCR with gold and silica particles of comparable size and charge, without changing the typical PCR conditions such as reagent concentrations and amplification profile.To evaluate the overall ability of AuNPs to enhance PCR in terms of specificity and yield (efficiency), two PCR models were examined: (1) nitrogen fixation (NifD) gene from A. brasilense Sp7 bacterial culture (A. brasilense DNA detection, 710 bp amplicon), and (2) chromosomal ompA gene of C. trachomatis (157 to 160 bp target sequences for the variable domain 2 of MOMP). For brevity, we will refer to PCR models 1 and 2, respectively.
First, an optimal concentration of 16 nm citrate-stabilized AuNPs was found to be 0.4 nM by adding AuNPs in various concentrations to both PCR systems (Fig. S3, ESI†). This result is in good agreement with previous reports for quite different PCR models.29,32Fig. 2a illustrates the specificity of AuNP-assisted PCR performed with model 1. Numerous nonspecific bands (smears) are observed for duplicate lanes 1 and 2 without nanoparticles, performed by using DNA template mixed with heterogeneous DNA fragments from similar microorganisms, whereas the addition of citrate-stabilized AuNPs at an optimal 0.4 nM concentration completely inhibits all non-target bands. Fig. 2b exemplifies the efficiency of AuNP-assisted PCR, when no significant bands can be observed without nanoparticles (duplicate lanes 1 and 2) in the case of a low DNA template concentration, performed with ten-fold diluted initial template DNA. It should be emphasized, that even for this unoptimized system, the addition of AuNPs at optimal 0.4 nM concentration results in a great PCR yield enhancement. To the best of our knowledge, this is the first demonstration of the yield-enhanced AuNP-PCR for A. brasilense Sp7 (NifD) gene. Additionally, we have observed the enhancing effect of AuNPs on two-round error-prone PCR system by taking the PCR product as a template for the second round of PCR amplification (Fig. S5, ESI†). Moreover, we have shown that AuNPs could be used to improve PCR specificity while amplifying the long PCR product (Fig. S6, ESI†) which is in good agreement with a recent study by Zhou et al.33 Finally, in the third test with the model 1 (Fig. 2c), the AuNPs were added after amplification step to verify whether the AuNPs affect the amplification step themselves (as in Fig. 2a and b) or the post-amplification PCR outcome, just before gel electrophoresis. As shown on Fig. 2c, there was no effect of AuNPs added after amplification step without nanoparticles and the smeared nonspecific bands look similarly for lanes 1 and 2. This gives strong evidence for the first assumption – citrate-stabilized AuNPs enhance the PCR outcome just during the amplification step.
One of hypothetic mechanisms of nano-PCR29 is that AuNPs exhibit SSB-like behavior related to greater affinity of single-stranded DNA-binding protein for ssDNA compared to dsDNA. However, this explanation remains unanswered the following question: how many AuNPs per one PCR tube are needed to perform efficient PCR amplification caused by an SSB-like moiety alone? Below we provide simple estimations based on our experimental conditions. A typical PCR mixture, used in our experiments with optimal concentration 0.4 nM of AuNPs,29 contained 2.5 U of Taq DNA-polymerase, equal to 10 ng or 3.5 nM of enzyme (the activity of 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 U Taq-DNA polymerase is equal to 4 ng or 43 fmol of 94 kDa Mw protein). A rough estimation shows that the molar ratio between Taq DNA-polymerase and AuNPs is about 1
000 U Taq-DNA polymerase is equal to 4 ng or 43 fmol of 94 kDa Mw protein). A rough estimation shows that the molar ratio between Taq DNA-polymerase and AuNPs is about 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10. The number of DNA templates, which exponentially grows in each of 35 PCR cycles, is at least one order greater than the number of DNA-polymerase molecules per one PCR tube. This means that the molar ratio DNA
10. The number of DNA templates, which exponentially grows in each of 35 PCR cycles, is at least one order greater than the number of DNA-polymerase molecules per one PCR tube. This means that the molar ratio DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) AuNPs is about 1
AuNPs is about 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, which is quite insufficient to ensure AuNP action as an SSB-like substances.34 Consequently SSB-like behavior of gold nanoparticles seems to be not relevant to AuNP-assisted PCR enhancement.
1, which is quite insufficient to ensure AuNP action as an SSB-like substances.34 Consequently SSB-like behavior of gold nanoparticles seems to be not relevant to AuNP-assisted PCR enhancement.
To investigate a possible role of surface functionalization, we used AuNPs stabilized with two neutral polymer ligands such as PVP (AuNPs–PVP) and mPEG-SH (AuNPs–PEG-SH). According to Table 1 data, the both polymers do not change the sign and, in some extent, the value of nanoparticle zeta-potential. Surprisingly enough, the enhancing effect of AuNPs–PVP and AuNPs–PEG-SH on PCR was negligible (Fig. 3).
In particular, non-specific bands were present even at high concentrations of nanoparticles, and the PCR tubes remained pink colored after amplification (Fig. 4). In some cases, the addition of a high nanoparticle amount resulted in complete inhibition of PCR (see, e.g., lanes 11 and 7 in Fig. 3a and b, respectively). We note that all these experiments included reagent controls with free polymers added to PCR tubes to verify the polymer effect alone (see lanes SA in Fig. 3). Thus, the functionalization of AuNP surface with neutral polymers, which does not change the initial negative charge of AuNPs, results in complete suppression of the PCR enhancement. This observation can be attributed to the steric hindrance for biomolecular PCR components by protective PVP or mPEG-SH layers. In particular, these stabilizers prevent the nanoparticle aggregation after PCR amplification (Fig. 4b). By contrast, the stabilization of AuNPs with small citrate ions results in great enhancing PCR outcome (Fig. 2a and b), but does not prevent nanoparticle aggregation at the end of PCR (Fig. 4a).
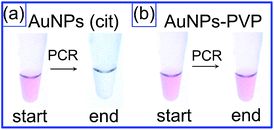 | ||
| Fig. 4 Photos of test-tubes with citrate- (a) and PVP-stabilized (b) AuNPs before and after PCR amplification. | ||
The next set of experiments was aimed at understanding the role of AuNP functionalization with positively charged ligands. These experiments were motivated by a recent study,30 revealing a crucial role of the particle charge in PCR enhancing. Specifically, the optimal enhancing concentration of positively-charged PDDA-capped AuNPs was shown30 to be 1.54 pM, which is 260 times lower than the optimal 400 pM concentration of negatively charged citrate-stabilized AuNPs.29 The goal of our experiment was twofold: (1) to verify the enhancing role of PDDA ligand for PCR models that differs from those investigated in ref. 30; (2) to examine the enhancing role of positive nanoparticle charge with CTAB ligand, which differs from PDDA. Clearly, the affirmative results for both points would expand the reported observations30 and confirm a universal role of the positive AuNP charge in PCR enhancing. However, our experiments with models 1 and 2 gave negative answers for both questions. Fig. 5a demonstrates the presence of non-target bands for very small (0.001, 0.01 pM) and high (10, 100 pM) concentrations of positively charged CTAB-stabilized AuNPs. Furthermore, for the same experimental model, no significant enhancement of PCR outcome was observed with PDDA-capped AuNPs (Fig. 5b) at variation of nanoparticle concentration from 0.001 pM to 5 pM. Finally, when we used the positively charged CTAB-stabilized gold nanorods, the results were the same: no PCR enhancing or even complete inhibition. This illustrates insignificant role of the nanoparticle shape, at least for the PCR models examined here. Similar negative results were obtained for second PCR model, i.e. with nitrogen fixation (NifD) gene from A. brasilense Sp7 (Fig. S4, ESI†). Thus, our experiments with two PCR models and with two cationic polymers did not confirm any efficient PCR enhancing reported previously30 for a different PCR model and positively charged AuNPs with PDDA ligand.
The precise reasons for observed discrepancy between our results and previously reported ones are unclear at present. Furthermore, the observed effects at trace 0.001 AuNP concentrations cannot be explained in terms of existing mechanistic models. In any case, we can conclude that the enhancing role of positively charged AuNPs is not ubiquitous. Instead, it can be ligand-dependent and PCR model-dependent.
Another proposed mechanism of AuNP-assisted PCR is related to high thermal conductivity of AuNPs as compared to water.32 This can change the amplification conditions in local environment around gold nanoparticles, thus changing the local conditions in “nanoreactor”. On the other hand, it has been supposed30 that Au nanoparticles in PCR mixture act as small nanoreactors, in which the local concentration of primers and templates is increased owing to electrostatic interactions between AuNPs and PCR components. In this model, the particle charge is important, irrespective of the particle material. To verify these models, we fabricated silica nanospheres with the 20 nm size and −31.2 mV zeta-potential close to those for citrate-stabilized AuNPs (16 nm and −29.9 mV, respectively). If the nanoparticle charge plays a crucial role, one would expect a similar PCR enhancement for both particle types. If the material heat properties are important, one would expect quite different PCR outcomes. Fig. 6 shows that the second alternative agrees with experimental data. Indeed, we observed dramatic difference in PCR results for gold and silica nanoparticles with close parameters except for their heat properties. Of course, the chemical structure and physical properties of the nanoparticle surface should also be considered for ultimate conclusions.
3. Conclusion
In conclusion, we have studied the effects of AuNP concentration, charge, shape and material on the PCR efficiency and specificity. For two new PCR systems that were not studied previously, we have obtained high specific and efficient enhancement of target amplicons at optimal citrate-stabilized AuNP concentration about 0.4 nM. This observation agrees with previous reports for other PCR models and confirms a universal nature of underlying enhancement mechanisms at least for citrate-stabilized AuNPs.29,32–36 Functionalization of citrate-stabilized AuNPs with neutral PVP and PEG-SH polymers results in elimination of enhancing properties, although the particle zeta-potentials remain the original sign and values. This phenomenon can be related to steric hindrance effects that prevent a proper interaction of PCR molecular components with nanoparticles. Functionalization of AuNPs with cationic polymers PDDA and CTAB leads to expected reverse of the particle charge sign. However, in contrast to previous report on positive enhancing properties of PDDA capped particles, we have obtained non-distinguishable enhancing effect. The particle shape plays negligible role as demonstrated by comparison of PCR outcomes with CTAB-coated gold nanospheres and nanorods. We have performed also PCR amplifications with two gold and silica particles of similar size and charge to show that the gold particles greatly enhance PCR results whereas silica counterparts do not. This implies a possible positive role of metallic heat properties in local environment of PCR nanoreactor. To summarize, further studies are needed to understand the underlying physicochemical mechanisms of nanoparticle-enhanced PCR, including the role of nanoparticle size, charge, surface functionalization, and the dynamic ratio between nanoparticle concentration and concentration of PCR system components.4. Experimental section
4.1 Materials
All chemicals were obtained from commercial suppliers and used without further purification. Cetyltrimethylammonium bromide (CTAB, >98.0%), L-ascorbic acid (AA, >99.9%), sodium borohydride (NaBH4, 99%), tetraethyl orthosilicate (TEOS), aqua ammonia (28–30%), L-arginine (98%), cyclohexane (anhydrous, 99.5%), thiolated poly(ethylene glycol) (mPEG-SH, Mw = 5000), poly(vinyl pyrrolidone) (PVP, Mw = 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000), polydiallyldimethylammonium chloride (PDDA, 20% solution) were purchased from Sigma-Aldrich. Hydrogen tetrachloroaurate trihydrate (HAuCl4·3H2O) and silver nitrate (AgNO3, >99%) were procured from Alfa Aesar. Absolute ethanol (ET0016, 99.99%) was obtained from Scharlau Chemie. Sodium hydroxide (NaOH), potassium carbonate (K2CO3) and sodium chloride (NaCl) were purchased from Reachim (Russia). Hydrochloric acid (HCl, 37 wt% in water) was purchased from Vekton (Russia). Tris, (C4H11NO3); ethylenediaminetetraacetic acid, disodium salt (EDTA); boric acid (HBO3); LE-2 agarose (Biotechnology Grade), ethidium bromide (EtBr), were purchased from Helicon (Russia). Phosphate buffer in tablets (PB; pH 7.4), 0.01 Mwas purchased from Biolot. MassRuler Low Range DNA Ladder, 6× loading dye, Taq DNA polymerase, dNTP mix were purchased from Fermentas, ThermoFisher Scientific. Ultrapure water was obtained from a Milli-Q Integral 5 system (Millipore) and used in all experiments.
000), polydiallyldimethylammonium chloride (PDDA, 20% solution) were purchased from Sigma-Aldrich. Hydrogen tetrachloroaurate trihydrate (HAuCl4·3H2O) and silver nitrate (AgNO3, >99%) were procured from Alfa Aesar. Absolute ethanol (ET0016, 99.99%) was obtained from Scharlau Chemie. Sodium hydroxide (NaOH), potassium carbonate (K2CO3) and sodium chloride (NaCl) were purchased from Reachim (Russia). Hydrochloric acid (HCl, 37 wt% in water) was purchased from Vekton (Russia). Tris, (C4H11NO3); ethylenediaminetetraacetic acid, disodium salt (EDTA); boric acid (HBO3); LE-2 agarose (Biotechnology Grade), ethidium bromide (EtBr), were purchased from Helicon (Russia). Phosphate buffer in tablets (PB; pH 7.4), 0.01 Mwas purchased from Biolot. MassRuler Low Range DNA Ladder, 6× loading dye, Taq DNA polymerase, dNTP mix were purchased from Fermentas, ThermoFisher Scientific. Ultrapure water was obtained from a Milli-Q Integral 5 system (Millipore) and used in all experiments.
4.2 Synthesis of nanoparticles
Spherical AuNPs with 16 nm average diameter were fabricated by reduction of HAuCl4 with sodium citrate as described by Grabar et al.37 and were used as a template for subsequent capping with various surface ligands. Briefly, 25 mL of 38.8 mM sodium citrate was added quickly to boiled 250 mL of 1 mM water solution HAuCl4, which resulted in a change in solution color from pale yellow to deep red.To obtain polymer-capped AuNPs,38 the as-prepared AuNPs were mixed with appropriate amount of each surface agent such as mPEG-SH, PVP or PDDA respectively, incubated for some time with subsequent centrifugation and resuspension in MQ water. A detail description of ligand capping procedures are given in ESI file.†
Gold nanorods (AuNRs) with length and diameter about 70 nm and 25 nm, respectively, were fabricated according to Ratto et al.39 with minor modifications.40 First, gold seed particles were prepared by adding aqueous ice-cold sodium borohydride (10 mM, 0.1 mL) to a mixed aqueous solution of CTAB (0.1 M, 1 mL) and HAuCl4 (10 mM, 0.025 mL) and were vigorously stirred for about 2 minutes. After sequential addition of 28 μL of 100 mM AA and 12 μL of two-hours-aged gold seeds to a growth solution (5 mL of 0.1 M CTAB, 250 μL of 0.01 M HAuCl4, and 100 μL of 4 mM AgNO3), the mixture was incubated for 24 h at 25 °C. Finally, 16 μL of 10 mM ascorbic acid was added in three portions every 24 h. AuNRs were allowed to grow overnight without stirring at 30 °C. Then, as-prepared AuNRs were repeatedly centrifuged and redispersed overnight in 1 mM CTAB.
Highly monodisperse SiO2NPs with 20 nm TEM average diameter were synthesized following the multistep seed-mediated growth technique in an aqueous solution of L-arginine.41 Briefly, 9.1 mg of L-arginine was added to 6.9 mL of water in a standard 20 mL scintillation vial under magnetic stirring. Then, 0.45 mL of cyclohexane was accurately added to the top of the solution and the mixture was heated to 60 °C. Further, 0.55 mL of TEOS was added to the top layer of cyclohexane and the mixture was allowed to react for 20 h. A key tip for synthesizing high-quality monodisperse SiO2NPs is to keep the cyclohexane and water parts unmixed, ensuring very slow addition of TEOS to the reaction mixture.
4.3 Characterization
Extinction spectra of as-prepared and ligand-capped nanoparticles solutions were measured with a Specord BS-250 and Specord S-300 spectrophotometers (Analytik, Jena, Germany). Transmission electron microscopy (TEM) images were obtained using Libra-120 transmission electron microscope (Carl Zeiss, Jena, Germany) at the Simbioz Center for the Collective Use of Research Equipment in the Field of Physical–Chemical Biology and Nanobiotechnology at the IBPPM RAS. The average diameters of as-prepared and polymer-capped AuNPs and silica nanoparticles and their zeta-potentials were measured with a Zetasizer Nano ZS device (Malvern, UK).4.4 DNA samples
| Primer name | Sequence (5′–3′) | Length (bp) | Amplicon size (bp) | Ref. |
|---|---|---|---|---|
| Primers used for A. brasilense Sp7 DNA | ||||
| nifD-up | GAGGCTATTCGGCTATGACTG | 21 | 710 (sample 1) | 43 |
| nifD-do | CGAGATCATCGCCGTCGGGC | 20 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Primers used for C. trachomatis DNA | ||||
| momp-fw1 | TTCAATTTAGTTGGATTGTTTGG | 23 | 157–160 (sample 2) | 45 |
| momp-fw2 | TCAACTTAGTTGGCTTATTCGG | 22 | ||
| momp-fw3 | TCAATTTAGTGGGGTTATTCGG | 22 | ||
| momp-rv1 | CACATTCCCAGAGAGCTGC | 19 | ||
| momp-rv2 | CACATTCCCACAAAGCTGC | 19 | ||
| momp-rv3 | CGGACTCCCACAAAGCTGC | 19 | ||
| momp-rv4 | GCACTCCCACAAAGCTGC | 18 | ||
| F1 | CGGTATTAGTATTTGCCGCTTTG | 23 | 1156 (sample 3) | 46 |
| B11 | CGGAATTGTGCATTTACGTGAG | 22 | ||
4.5 PCR and gel electrophoresis
All PCR reactions were conducted in triplicate with a T-100 Thermal Cycler (Bio-Rad, USA). PCR reagents were mixed to a final 30 μL reaction volume in 200 μL thin-walled tubes (Scientific Specialties, Inc., USA). Full PCR mixture in absence of nanoparticles was used as blank in all experiments. Pure double-distilled water (PCR-grade) as template was used for negative control in all experiments.Each PCR tube with sample 1 DNA (A. brasilense Sp7) as a template contained: 3 μL 10× PCR buffer, 0.6 μL 10 mM dNTPs, 3 μL 25 mM MgCl2, 0.5 μL 5 U μL−1 Taq DNA polymerase, 2 μL template containing 120–200 ng DNA, 50 pmol of each primer (0.3 μL nifD-up and 0.3 μL nifD-do primer), and 15 μL of appropriate NPs (varying in concentrations and particle type) and filled up to the 30 μL final volume with MQ water. The PCR procedure was as follows: 5 min at 94 °C for pre-denaturation, followed by 35 cycles of 30 s at 94 °C, 30 s at 52 °C, and 30 s at 72 °C. Then the cycling was terminated after 10 min final elongation step at 72 °C.
Each PCR tube with sample 2 DNA (C. trachomatis) as a template contained: 3 μL 10× PCR buffer, 0.6 μL 10 mM dNTPs, 3 μL 25 mM MgCl2, 0.5 μL 5 U μL−1 Taq DNA polymerase, 2 μL template containing 120–200 ng DNA, a multiplex-broad-spectrum PCR primer mix containing 1 μL multiple forward primer (equimolar mix of 10 pmol momp-fw1, momp-fw2, momp-fw3) and 1.2 μL multiple reverse primer (equimolar mix of 10 pmol momp-rv1, momp-rv2, momp-rv3, momp-rv4), and 15 μL of appropriate NPs (varying in concentrations and particle type) and filled up to the 30 μL final volume with MQ water. The PCR procedure was as follows: 2 min preheating step at 94 °C followed by 38 cycles of amplification (30 s at 94 °C, 30 s at 55 °C, and 30 s at 72 °C) and a final 10 min elongation step at 72 °C. Purified DNA of genovar E as a template was used as positive control in all experiments.
Each PCR tube with sample 3 DNA (C. trachomatis) as a template contained: 3 μL 10× PCR buffer, 0.6 μL 10 mM dNTPs, 1.8 μL 25 mM MgCl2, 0.15 μL 5 U μL−1 Taq DNA polymerase, 1 μL template containing 120–200 ng DNA, 50 pmol of each primer (1 μL F1 and 1 μL B11), and 15 μL of AuNPs in serial dilutions and filled up to the 30 μL final volume with MQ water. The PCR procedure was as follows: 2 min preheating step at 92 °C followed by 35 cycles of amplification (45 s at 94 °C, 45 s at 55 °C, and 60 s at 72 °C) and a final 10 min elongation step at 72 °C. Purified DNA of genovar E as a template was used as positive control in all experiments.
After the PCR amplification, all the samples were stored at 4 °C before gel-electrophoresis. The PCR products were analyzed by horizontal electrophoresis system (SE 1, Helicon) with a voltage set at 125 V for 65 min (Elf-4 power supplier, DNA-technology) with EtBr staining. Briefly, 3.5 μL of each PCR product was mixed with 0.5 μL 6× loading dye prior to loading in the wells of an 1.5% agarose gel with EtBr containing in 1× TBE buffer (10 mM Tris, 10 mM boric acid, 1 mM EDTA, pH 8.0). The fluorescence of the bands was visualized by UV transillumination (312 nm, Vilber Lourmart), the gel electrophoregrams were obtained with a Canon 350D digital camera equipped with an orange filter to minimize the UV-lamp background lighting.
4.6 PCR product quantification
The performance of PCR additives was quantified according to recent report30 through calculations of two densitometric quantities, which are termed specificity and yield efficiency. The gel electrophoregram images were analyzed with an open ImageJ software and the output data were represented as the means and standard deviations (n = 3). A detail description of calculations is given in ESI file.† In brief, the ratio of the densitometric value of the specific target band to that of all bands amplified by PCR was defined as the specificity of amplification. By this definition, if there are no smears in PCR products, the maximal value of specificity equals 1. The ratio of the densitometric intensity of a specific target band to that for 500 bp of DNA marker was used as a measure of the yield efficiency. For brevity, we use the term “efficiency”. If the obtained efficiency is ≥1, the PCR additive is considered as high efficient for PCR optimization. In all figures, molecular weight markers (M) represented the same SM0383 (ThermoFisher Scientific). The optimal concentration of PCR additives was defined as a concentration that maximizes the brightest of specific target band.Acknowledgements
This work was supported the Russian Scientific Foundation (project No. 14-13-01167). The work by TEP (PCR amplification) was partially supported by Bortnik Foundation grant U. M. N. I. K. (project No. 11736 p/17259). The work by VAK (synthesis of silica nanoparticles) was partially supported by RFBR grant (project No. 16 52-45026). We thank Drs V. A. Bogatyrev, L. A. Dykman and B. N. Khlebtsov for helpful discussion, Dr A. M. Burov for TEM experiments, A. A. Budanova and Dr A. A. Shirokov for providing the Azospirillum brasilense Sp7 cells and help in the DNA extraction experiments, Dr T. I. Polyanina and Yu. Saltykov for providing the specimen from clinical samples.Notes and references
- R. K. Saiki, D. H. Gelfand, S. Stoffel, S. J. Scharf, R. Higuchi, G. T. Horn, K. B. Mullis and H. A. Erlich, Science, 1988, 239, 487 CAS.
- J. Lu, H. Li, H. An, G. Wang, Y. Wang, M. Li, Y. Zhang and J. Hu, J. Am. Chem. Soc., 2004, 126, 11136 CrossRef PubMed.
- C. Ding and C. R. Cantor, Proc. Natl. Acad. Sci. U. S. A., 2003, 100, 7449 CrossRef CAS PubMed.
- J. Sambrook and D. W. Russell, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 3rd edn, 2001 Search PubMed.
- W. Henke, K. Herdel, K. Jung, D. Schnorr and S. A. Loening, Nucleic Acids Res., 1997, 25, 3957 CrossRef CAS PubMed.
- M. Nagai, A. Yoshida and N. Sato, Biochem. Mol. Biol. Int., 1998, 44, 157 CAS.
- P. Winship, Nucleic Acids Res., 1989, 17, 1266 CrossRef CAS PubMed.
- Q. Chou, Nucleic Acids Res., 1992, 20, 4371 CrossRef CAS PubMed.
- P. Belgrader, W. Benett, D. Hadley, G. Long, R. Mariella Jr, F. Milanovich and S. Nasarabadi, et al. , Clin. Chem., 1998, 44, 2191–2194 CAS.
- C. T. Wittwer, K. M. Ririe, R. V. Andrew, D. A. David, R. A. Gundry and U. J. Balis, BioTechniques, 1997, 22, 176–181 CAS.
- K. H. Roux, Genome Res., 1995, 4, S185–S194 CrossRef CAS.
- S. Ekman, Lichenologist, 1999, 31, 517–531 CrossRef.
- K. H. Hecker and K. H. Roux, BioTechniques, 1996, 20, 478–485 CAS.
- R. T. D'Aquila, L. J. Bechtel, J. J. Videler, J. J. Eron, P. Gorczyca and L. C. Kaplan, Nucleic Acids Res., 1991, 19, 3749 CrossRef.
- H. A. Erlich, D. Gelfand and J. J. Sninsky, Science, 1991, 252, 1643–1651 CrossRef CAS PubMed.
- M. Rabodonirina, L. Cotte, A. Boibieux, K. Kaiser, M. Mayencon, D. Raffenot, C. Trepo, D. Peyramond and S. Picot, J. Clin. Microbiol., 1999, 37, 127 CAS.
- R. Don, P. Cox, B. Wainwright, K. Baker and J. Mattick, Nucleic Acids Res., 1991, 19, 4008 CrossRef CAS PubMed.
- X. Lou and Y. Zhang, ACS Appl. Mater. Interfaces, 2013, 5, 6276–6284 CAS.
- J. Jia, L. Sun, N. Hu, G. Huang and J. Weng, Small, 2012, 8, 2011–2015 CrossRef CAS PubMed.
- L. Ma, S. He, J. Huang, L. Cao, F. Yang and L. Li, Biochimie, 2009, 91, 969–973 CrossRef CAS PubMed.
- S.-H. Hwang, S.-G. Im, S. S. Hah, V. T. Cong, E. J. Lee, Y.-S. Lee, G. K. Lee, D.-H. Lee and S. J. Son, PLoS One, 2013, 8, e73408 CAS.
- Y. Liang, F. Luo, Y. Lin, Q. Zhou and G. Jiang, Carbon, 2009, 47, 1457–1465 CrossRef CAS.
- R. M. Williams, S. Nayeem, B. D. Dolash and L. J. Sooter, PLoS One, 2014, 9, e94117 Search PubMed.
- Z. Z. Zhang, M. C. Wang and H. J. An, Nanotechnology, 2007, 18, 355706 CrossRef.
- H. B. Shen, M. Hu, Y. B. Wang and H. Q. Zhou, Biophys. Chem., 2005, 115, 63–66 CrossRef CAS PubMed.
- L. Wang, Y. Zhu, Y. Jiang, R. Qiao, S. Zhu, W. Chen and C. Xu, J. Phys. Chem. B, 2009, 113, 7637–7641 CrossRef CAS PubMed.
- A. Khaliq, P. J. Sonawane, B. K. Sasi, B. S. Sahu, T. Pradeep, S. K. Das and N. R. Mahapatra, Nanotechnology, 2010, 21, 255704 CrossRef PubMed.
- W. X. Y. Cao, X. Y. Shi, W. C. Yang, X. D. Zhang, C. H. Fan and J. Hu, Analyst, 2009, 134, 87–92 RSC.
- H. Li, J. Huang, J. Lv, H. An, X. Zhang, Z. Zhang, C. Fan and J. Hu, Angew. Chem., 2005, 44, 5100–5103 CrossRef CAS PubMed.
- L. Yuan and Y. He, Analyst, 2013, 138, 539–545 RSC.
- D. Cui, F. Tian, Y. Kong, I. Titushikin and H. Gao, Nanotechnology, 2004, 15, 154–157 CrossRef CAS.
- M. Li, Y.-C. Lin and C.-C. Wu, Nucleic Acids Res., 2005, 33, e184 CrossRef PubMed.
- A. Li, B. Zhou, C. S. Alves, B. Xu, R. Guo, X. Shi and X. Cao, ACS Appl. Mater. Interfaces, 2016, 8, 25808–25817 CAS.
- S. Dabrowski, M. Olszewski, R. Piatek, A. Brillowska-Dabrowska, G. Konopa and J. Kur, Microbiology, 2002, 148, 3307–3315 CrossRef CAS PubMed.
- A. L. Haber, K. R. Griffiths, A. K. Jamting and K. R. Emslie, Anal. Bioanal. Chem., 2008, 392, 887–896 CrossRef CAS PubMed.
- B. V. Vu, D. Litvinov and R. C. Willson, Anal. Chem., 2008, 80, 5462–5467 CrossRef CAS PubMed.
- K. C. Grabar, R. G. Freeman, M. B. Hommer and M. J. Natan, Anal. Chem., 1995, 67, 735–743 CrossRef CAS.
- K. Aslan and V. H. Pérez-Luna, Langmuir, 2002, 18, 6059–6065 CrossRef CAS.
- F. Ratto, P. Matteini, F. Rossi and R. Pini, J. Nanopart. Res., 2010, 12, 2029 CrossRef CAS.
- T. E. Pylaev, V. A. Khanadeev, B. N. Khlebtsov, L. A. Dykman, V. A. Bogatyrev and N. G. Khlebtsov, Nanotechnology, 2011, 22, 285501 CrossRef CAS PubMed.
- K. D. Hartlen, A. P. T. Athanasopoulos and V. Kitaev, Langmuir, 2008, 24, 1714–1720 CrossRef CAS PubMed.
- http://collection.ibppm.ru/ .
- D. P. Potrich, L. M. Passaglia and I. S. Schrank, Braz. J. Med. Biol. Res., 2001, 34, 1105–1113 CrossRef CAS PubMed.
- J. Döbereiner and J. M. Day in Intern. Symp. Nitrogen Fixation – Interdisciplinary Discussion, 1976, pp. 518–538 Search PubMed.
- K. D. Quint, L. J. Van Doorn, B. Kleter, M. N. C. De Koning, H. A. M. Van Den Munckhof, S. A. Morre, B. Ter Harmsel, E. Weiderpass, G. Harbers, W. J. G. Melchers and W. G. V. Quint, J. Mol. Diagn., 2007, 9, 631–638 CrossRef CAS PubMed.
- D. Dean, E. Oudens, G. Bolan, N. Padian and J. Schachter, J. Infect. Dis., 1995, 172, 1013–1022 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: Extinction spectra of as-prepared gold and silica nanoparticles; extinction spectra and photos of as-prepared and functionalized NPs before and after addition of NaCl salt; chemical structures and IUPAC names of surface ligands; evaluation of nano-PCR specificity and efficiency; the effect of AuNPs on PCR amplification of 710 bp target from A. brasilense and a 160 bp target from C. trachomatis, the effect of AuNPs–CTAB, AuNPs–PDDA and AuNRs-662 on PCR model 1; the effect of AuNPs on two-round PCR model 1; the effect of AuNPs on PCR amplification of a long 1156 bp target from PCR model 3. See DOI: 10.1039/c6ra20472d |
| This journal is © The Royal Society of Chemistry 2016 |




