Facile one-step targeted immobilization of an enzyme based on silane emulsion self-assembled molecularly imprinted polymers for visual sensors†
Guoning
Chen
,
Hua
Shu
,
Lu
Wang
,
Kamran
Bashir
,
Qun
Wang
,
Xia
Cui
,
Xuan
Li
,
Zhimin
Luo
,
Chun
Chang
and
Qiang
Fu
 *
*
School of Pharmacy, Xi'an Jiaotong University, Xi'an 710061, China. E-mail: fuqiang@mail.xjtu.edu.cn
First published on 5th November 2019
Abstract
Immobilized enzymes play significant roles in many practical applications. However, the enzymes need to be purified before immobilization by conventional immobilizing methods, and the purification process is expensive, laborious, complicated and results in a decrease of the enzymatic activity. So, we present a novel method by a facile one-step targeted immobilization of an enzyme without a purification process from complex samples. For this purpose, a novel molecularly imprinted polymer was prepared via a silane emulsion self-assembly method using boric acid-modified Fe3O4 nanoparticles as magnetic nuclei, horseradish peroxidase as a template, 3-aminopropyltriethoxysilane as a functional monomer and tetraethyl orthosilicate as a crosslinking agent. The molecularly imprinted polymers were characterized using a scanning electron microscope, X-ray photoelectron spectroscope, vibrating sample magnetometer and X-ray diffractometer. The as-prepared and characterized materials were employed to immobilize horseradish peroxidase from a crude extract of horseradish. Moreover, the immobilized horseradish peroxidase was employed to develop visual sensors for the detection of glucose and sarcosine. This study demonstrated that the molecularly imprinted polymers prepared via the silane emulsion self-assembly method can facilely immobilize horseradish peroxidase from a crude extract of horseradish without any purification process. The developed visual method based on the immobilized horseradish peroxidase shows great potential applications for the visual detection of glucose and sarcosine.
Introduction
Enzymes are significant biological catalysts that have been widely used in food safety,1 environmental monitoring2 and clinical diagnosis3 owing to their good selectivity, high activity, mild reaction conditions and fast detection. It is well known that the use of enzymes is limited by the realistic difficulties of separation and instability. Enzyme immobilization is a straightforward method to overcome these problems, which fixes the enzymes onto various selected supports. Conventional methods for immobilizing enzymes are mainly divided into four categories: adsorption,4 covalent binding,5 entrapment6 and cross-linking.7 However, it is necessary for the target enzymes to be purified before immobilization using conventional immobilizing methods. The high cost for the purification, the complex purification process and the loss in enzymatic activity are often cited as problems. Thus, the development of a directly immobilized method for target enzymes from complex samples without purifying is highly desired.To the best of our knowledge, a directly immobilized method for target enzymes from complex samples without purification has rarely been reported. Antibodies, aptamers and molecularly imprinted polymers (MIPs) can be used to selectively identify target molecules from complex samples. Compared with antibodies and aptamers, MIPs are considered promising carriers for specifically immobilizing target enzymes because of their good stability, low-cost, short development time and easy production.8 The effectiveness of molecular imprinting for small molecules has been well demonstrated. However, the imprinting of biological macromolecules (such as proteins) is challenging because of their properties, such as a large molecular size, water solubility, high flexibility and complex surface structure.9 In recent years, to overcome the aforementioned drawbacks, scientists have proposed a variety of approaches, including surface imprinting,10 epitope imprinting,11 and Pickering emulsion imprinting12 and so on.13
Many enzymes have been proved to be glycoproteins. Recently, using boronic acids as ligands to synthesize MIPs for the specific recognition of glycoproteins has demonstrated highly applicable values.14–18 Boronate affinity molecularly imprinted polymers can be produced by two steps: (1) boric acid groups were grafted onto selected carriers and their covalent binding with the target glycoprotein, and (2) construct imprinting layers on the surface of the complex by the surface imprinting method and remove the glycoprotein. The imprinted layers are usually constructed by the dopamine self-polymerization method, aniline polymerization method, sol–gel method and so on. As compared to other methods, the sol–gel method has the great advantages of superior physical rigidity, chemical inertness, thermal stability and hydrophilic properties, successfully applied in the preparation of imprinted polymers of small molecular compounds.19,20 However, some organic silane used to prepare the imprinted layers could only be dissolved in organic solvents and polymerized under catalysis by acid or alkali, which restrict the application of the sol–gel method in preparing protein MIPs. To overcome these limitations, a silane emulsion self-assembly reaction system without organic solvents or acid/base catalysts was designed and the imprinted polymer coating was formed in the presence of proteins.
Horseradish peroxidase (HRP), extracted from the roots of horseradishes, is widely used in analytical applications. HRP can effectively initiate the oxidation reaction of the peroxidase substrate of 3,3′,5,5′-tetramethylbenzidine (TMB) in the presence of H2O2, accompanied with a color change from colorless to blue. This reaction system has been employed to detect H2O2 molecules, or one can produce H2O2 molecules.21,22 Sarcosine and glucose are very important small biomolecules in body fluids. Sarcosine is related to the diagnosis of cancer of the prostate gland.23 The glucose level in blood is most commonly employed as a clinical indicator for diagnosing diabetes.24 Sarcosine and glucose cause a rapid release of H2O2 by the catalytic action of their oxidases (sarcosine oxidase and glucose oxidase).25,26 Thus, sarcosine and glucose can be detected through a visual method based on the catalytic reaction system described above.
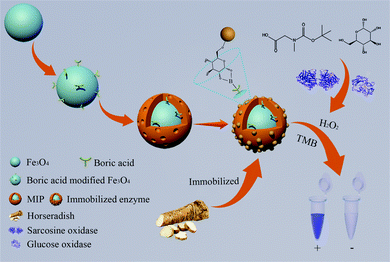
In this study, we present a new method for preparing carriers of immobilizing HRP based on the silane emulsion self-assembled boronate affinity molecularly imprinted polymers. The resultant products were used to immobilize HRP by a facile one-step process from a crude extract of horseradish without a purification process, and then the immobilized HRP was further developed as a visual sensor for detecting glucose and sarcosine. The principle and procedure of this work is shown in Fig. 1. The amino-functionalized Fe3O4 nanospheres were synthesized as a support for rapid magnetic separation. Boronic acid functionalized Fe3O4 nanoparticles were further synthesized to covalently bind with glycoprotein. Then, an imprinted shell was grafted onto the surface of the aforementioned carriers by a silane emulsion self-assembly method, in which 3-aminopropyltriethoxysilane (APTES) and tetraethyl orthosilicate (TEOS) were used as functional monomers and crosslinkers, respectively, and an aqueous solution containing tween-20 as a reaction system. After removing the templates, the final products can specifically capture the templates of HRP from a crude extract of horseradish, avoiding the complicated purification process of enzymes. The immobilized HRP also shows great potential in the visual detection of sarcosine and glucose.
Results and discussion
Preparation of MIPs
In this study, amino-functionalized Fe3O4 nanospheres were synthesized via a one-pot process to provide a magnetic response, making the obtained products separate conveniently. Then, FPBA was grafted onto the surface of the amino-functionalized Fe3O4 nanoparticles to introduce boronic acid groups that can covalently bind with the glycoprotein. Finally, an imprinted shell was grafted onto the surface of the aforementioned boric acid-modified Fe3O4 nanoparticles by a silane emulsion self-assembly method.The sol–gel method is a common method employed for preparing polymer layers of MIPs owing to its high chemical stability, moderate cost and easy preparation. However, some organic silanes could only be dissolved in organic solvents and polymerized under the catalyzing action of acid or alkali, which restrict the application of the sol–gel method in preparing protein MIPs. Based on this reason, this article described a silane emulsion self-assembly method, in which polymerization occurred spontaneously in an aqueous solution without acid or base catalysis. As shown in tube 2 of Fig. 2(a), the organic silane (APTES) acting as a functional monomer showed great miscibility with water. While another cross-linker monomer (TEOS) was not dissolved in water, and produced an immiscible two-phase fluid in the mixture. This immiscible solution was not polymerized. As shown in Fig. 2(b) and tube 1 of Fig. 2(a), the immiscible solution can be emulsified into a homogeneous emulsion in the presence of tween-20. Tween-20 is a surface-active agent and has no damage on the structure of the protein. This homogeneous emulsion brings about a rapid polymerization by self-reaction without any catalyst. The imprinting polymer layers were cloaked on the surface of the Fe3O4 nanoparticles covalently bound with the glycoprotein via the silane emulsion self-assembly method. After removing the templates, the boric acid affinity MIPs were obtained.
 | ||
| Fig. 2 The silane emulsion self-assembly method, (a) photographs of the polymerization reaction, (b) microscopic picture of the polymerization system. | ||
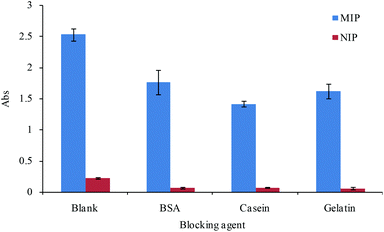
There is some non-specific binding of the protein to the developed MIPs because of abundant functional groups, such as –NH2 and –COOH, in the protein surface and some unavoidable non-specific binding sites in the MIP surface. In order to reduce the non-specific adsorption and improve the selectivity of the MIPs, non-specific adsorption groups were blocked by non-target proteins without affecting the specific interaction between the MIPs and the template. We tested some blocking agents, including BSA, casein and gelatin, in our experiments to reduce the non-specific binding. The effect of the blocking agent on the binding of the MIPs compared with the adsorption of NIPs was analyzed, and the results are shown in Fig. 3. The results showed that a similar blocking effect was obtained in the three different blocking agents, and the blocking agent of BSA was used in subsequent experiments owing to its better solubility.
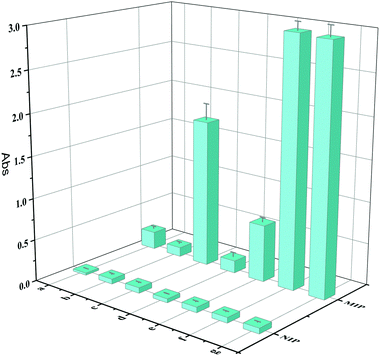
In order to obtain better target binding properties for the imprinted layers, some important parameters affecting the thickness and cavity numbers of the imprinting layers were optimized during the preparation procedure. The imprinting factor (IF), calculated according to the ratio of the enzyme activity of HRP captured by MIPs to that captured by NIPs,14 was used to evaluate the imprinting effect in this study. As shown in Fig. 4, the imprinting performance with different amounts of monomers ranging from 7.5 μL to 30 μL was investigated, and the optimal amount of TEOS was found to be 10 μL. An excessive amount of monomer led to the imprinting performance becoming dramatically worse; the reason for this was probably that the imprinting layer was too thick, and a thicker imprinting layer caused the burying of the recognition cavities. Too little monomer could not polymerize adequately with the template protein and form enough specific recognition cavities, and the non-specific recognition sites would be blocked with the blocking solution. The effect at different concentrations of HRP was then tested, and the best adsorption ability and IF value (45.08) were obtained at 500 μg mL−1 of HRP. When the HRP concentration was higher than 500 μg mL−1, the adsorption ability of the prepared MIPs was similar to that prepared at 500 μg mL−1, which may be attributed to the saturation of the template adsorbed on the carrier core of the boric acid-modified Fe3O4 nanoparticles. The adsorption of the MIPs decreased with decreasing concentration of HRP; the reason for this was that the imprinted cavity numbers decreased when the HRP concentration gradually decreased.
Characterization
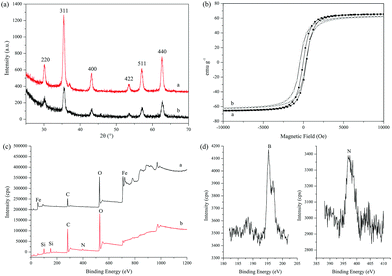
The structure details of the amino-functionalized Fe3O4 and MIPs were analyzed by an FT-IR spectrophotometer. As shown in Fig. S1(a),† the characteristic peak of the Fe–O group for the amino-functionalized Fe3O4 was observed at around 580 cm−1, and the absorption peaks at 3438 cm−1 and 1670 cm−1 were ascribed to the stretching and bending vibrations of N–H, indicating the successful synthesis of the amino-functionalized Fe3O4 nanoparticles. Compared with the amino-functionalized Fe3O4, a new peak of Si–O at 1050 cm−1 from the MIPs indicated that the SiO2-imprinted film was successfully coated onto the surface of the amino-functionalized Fe3O4.Fig. 5(a) shows the XRD diffractograms, and six peaks in 2θ at 30.20°, 35.40°, 43.14°, 53.56°, 57.02° and 62.70° with corresponding indices at (220), (311), (400), (422), (511) and (440), respectively, were shown, which matched well with the database of magnetite in the Joint Committee on Powder Diffraction Standards. The results showed that the prepared magnetic core was Fe3O4, and similar peaks in the diffractograms indicated that the synthesis process of MIPs did not change the crystal structure of Fe3O4.
VSM was used to characterize the magnetic properties of the amino-functionalized Fe3O4 nanospheres and MIPs. The magnetic hysteresis loops are shown in Fig. 5(b). The saturation magnetization values of the amino-functionalized Fe3O4 nanospheres and MIPs were 65.6 emu g−1 and 62.3 emu g−1, respectively. The imprinted polymer coating on the surface of the Fe3O4 nanospheres did not cause a significant decrease in the saturated magnetic strength of MIPs, and a high saturated magnetic strength of the MIPs will ensure rapid separation in the subsequent experiments. As shown in Fig. 5(b), there was hardly any magnetic hysteresis for the amino-functionalized Fe3O4 nanospheres and MIPs, indicating that they all showed superparamagnetic behavior.
The surface chemical element of the prepared nanoparticles was investigated by XPS, which was used to further confirm whether the imprinting layer was grafted successfully on the surface of the Fe3O4 nanoparticle. Fig. 5(c) (trace a) shows the XPS results of the boric acid-modified Fe3O4, and Fig. 5(d) shows the B and N signal peaks of the boric acid-modified Fe3O4. The signals of O, C, Fe, N and B appeared on the surface of the boric acid-modified Fe3O4. The XPS result of MIPs is shown in Fig. 5(c) (trace b). The signals of C, O N and Si are the main elements, and the visible signal of Si and the increased intensity of the N element were ascribed to the modified silicone emulsion self-assembly imprinted polymer. Compared with Fig. 5(c) trace a and Fig. 5(d), the emergence of the Si signal, increase of the N signal and disappearance of the Fe signal on the surface of the MIPs indicated that the imprinted layers were successfully introduced onto the surface of Fe3O4.
Fig. S2† shows the TGA curves of the amino-functionalized Fe3O4 and MIPs in the temperature range of 30–800 °C. As shown in Fig. S2,† the amino-functionalized Fe3O4 and MIPs were stable within 200 °C, and only about 2% weight was lost, which was owing to the loss of the absorbed moisture. The amino-functionalized Fe3O4 and MIPs continued to experience mass loss, and the temperature increased after 200 °C. The total mass losses of the amino-functionalized Fe3O4 and MIPs is 9% and 13%, respectively, and the additional mass losses from MIPs also indicated the surface imprinted polymer layers were constructed successfully.
The morphology of the prepared amino-functionalized Fe3O4 and MIPs was characterized using SEM and TEM. As shown in Fig. 6(a) and (c), the amino-functionalized Fe3O4 were spherical particles with an approximate size of 20 nm. After the imprinting process, microsphere particles (Fig. 6b) with a relatively larger size were obtained, indicating the deposition of the silicone emulsion self-assembly thin film over the surface of Fe3O4 successfully. As seen from Fig. 6d, it can be inferred that the thickness (5 nm) of the polymers were coated onto the surface of the amino-functionalized Fe3O4. It could be inferred that the thin imprinted films were successfully grafted onto the surface of amino-functionalized Fe3O4 from these images of SEM and TEM.
Selectivity
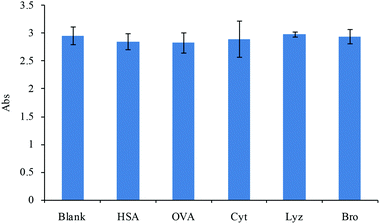
Selectivity is an important index, which determines the recognition performance of the developed MIPs in practical complex samples. The selectivity of MIPs for the substrate was investigated against the different interfering proteins, including HSA, OVA, Cyt, Lyz and Bro. The concentrations of the interfering proteins were the same as that of the substrate. The results are shown in Fig. 7. The MIPs exhibited a very high response intensity toward the target protein. In contrast, the additional interfering proteins showed almost no interference toward the recognition of the target protein. These results suggested that the MIPs exhibited excellent specificity toward HRP, even under the presence of interference of other protein.Activity assays
In this work, the enzyme catalysis reaction kinetics of the free and immobilized enzymes had been studied, and the value of Km had also been discussed. The results are shown in Fig. S3(a) and (c),† where the variation of the enzyme catalysis reaction kinetics of the free and immobilized enzyme showed a similar tendency. The rate is linear to the TMB concentrations at low concentration. However, the linear relationship deviates when the TMB concentrations increases. Moreover, Km was obtained from the Lineweaver–Burk plots of the double reciprocal of the Michaelis–Menten equation,28 1/ν = (Km/Vmax) × (1/[S]) + 1/Vmax, where ν is the initial velocity, Vmax is the maximal reaction velocity, Km is the Michaelis–Menten constant and [S] is the concentration of the substrate. The results are shown in Fig. S3(b) and (d);† the Km for this reaction of the free and immobilized enzymes obtained from this double reciprocal curve were 0.29 mM and 0.36 mM, respectively. However, the affinity of the immobilized enzyme to the substrate was slightly lower than that of the free enzyme, which may be ascribed to the space steric effect of MIPs.Immobilized horseradish peroxidase from horseradish
The direct immobilization of HRP from a crude extract of horseradish was used to evaluate the practical application of the MIPs. The principle and procedure are shown in Fig. 8(a). A target of HRP in the crude extract sample of horseradish is specifically captured by the developed MIPs. After washing the non-specific adsorbents, the HRP will be immobilized in three-dimensional holes of MIPs. Immobilized enzyme was detected by specific substrates. HRP was immobilized from a complex sample of horseradish directly by this procedure without extra steps of separation and purification. Thus, the extra time, high cost, resource waste and activity lost due to the separation and purification steps can be avoided. The result of enzyme immobilization is shown in Fig. 8(b). The signal produced by the immobilized enzyme of MIPs was significantly higher than that by the immobilized enzyme of NIPs. The reason is that the suitable hole for adsorption is insufficient in the surface of NIPs. It showed that HRP could be recognized and immobilized directly from a crude extract sample of horseradish, and other substances in the horseradish sample caused low interference in the process of immobilization. These results further confirm that the developed MIPs with extremely excellent selectivity have a great prospect in the application of directly immobilizing a target from an unpurified complex sample. | ||
| Fig. 8 The process (a) and result (b) diagrams of immobilizing HRP from a crude extract of horseradish. | ||
Reusability performance of MIPs
The reusability of the MIPs was evaluated by measuring their catalytic performance after the adsorption of HRP in consecutive adsorption–desorption cycles. The recovery derived from the ratio of the catalytic performance obtained at an arbitrary cycle to that at the first cycle was used to evaluate the reusability. Fig. S4† shows the results of the recovery. As can be seen, the MIPs only lost approximately 17.9% of their original catalytic performance after three adsorption–desorption cycles, and approximately 68.4% recovery of the original catalytic performance was still attained after five adsorption–desorption cycles. The above results indicated that the MIPs are stable and reusable.Analysis of glucose in plasma
HRP can catalyze the reaction of the peroxidase substrate 3,3,5,5,-tetramethylbenzidine (TMB) in the presence of H2O2, which has been employed to detect H2O2, or one can produce H2O2. The immobilized HRP showed similar catalytic activity to the HRP, so the immobilized HRP have practical applications in the detection for H2O2 and glucose because glucose causes a rapid release of H2O2 by the catalytic action of glucose oxidase. These days, a colorimetric method for the determination of glucose was established based on the immobilized enzyme combined with glucose oxidase. The results (Fig. 9(a)) suggested a good linear relation with the varied concentration of glucose (1–100 μg mL−1). The regression equation of a standard working curve is expressed as the following: A = 0.031c + 0.194, R2 = 0.9986. | ||
| Fig. 9 Standard curve for glucose (a) and sarcosine (c), and the selectivity of the developed method for glucose (b) and sarcosine (d) detection. | ||
The specificity of the method was investigated further to reveal the potential practicality, which was measured by assessing the response to other potential interfering chemicals including vitamin C, sucrose, glycine and mannitol. The results are shown in Fig. 9(b). These results clearly demonstrated that the interfering chemicals did not disturb the determination of glucose even though the concentrations were 10-fold higher than that of glucose. The results of this testing indicate that the method has high specificity toward glucose detection, which could be ascribed to the selectivity of glucose oxidase toward glucose.
The colorimetric method was finally applied to human serum in order to evaluate the applicability of the proposed method. The accuracy and precision of the method were evaluated at three concentration levels of spiked samples. The results are shown in Table 1; the accuracy, expressed as a percentage of recovery was within the range of 115.6–120.9%, and the relative standard deviations were less than 4.1%. These results indicate that the method had good accuracy and precision, which was enough validity of the proposed method for the quantitative determination of glucose in real samples. The proposed method was compared with other reported methods29–33 for the detection of glucose, and the results are listed in Table S1.† This method showed better selectivity and similar sensitivity.
| Spiked (μg mL−1) | Found (μg mL−1) | Recovery (%) | RSD (%) |
|---|---|---|---|
| 0 | 13.63 ± 0.56 | — | 4.1 |
| 5 | 19.68 ± 0.46 | 120.9 | 2.3 |
| 25 | 42.53 ± 0.99 | 115.6 | 2.3 |
| 50 | 74.04 ± 2.07 | 120.8 | 2.8 |
Analysis of sarcosine in urine
Immobilized HRP also showed practical application in the determination of sarcosine because H2O2 will be released rapidly when sarcosine is catalyzed by its own oxidase. Therefore, a colorimetric method for the determination of sarcosine was established based on the immobilized enzyme combined with sarcosine oxidase. The results (Fig. 9(c)) suggested a good linear relation with the varied concentration of sarcosine (0.1–10 μg mL−1). The regression equation of a standard working curve is expressed as the following: A = 0.0623c + 0.0347, R2 = 0.9985.The specificity of the method was also investigated and measured by assessing the response to other potential interfering chemicals, including glucose, vitamin C, glycine, phenylalanine and arginine. The results are shown in Fig. 9(d). These results clearly demonstrated that the interfering chemicals did not disturb the determination of sarcosine even though the concentrations were 10-fold higher than that of sarcosine. The results of this testing indicate that the method has high specificity toward sarcosine detection, which could be ascribed to the selectivity of sarcosine oxidase toward sarcosine.
The developed method was finally applied to detect sarcosine in urine to evaluate the applicability of the proposed method. The accuracy and precision of the method were evaluated at three concentration levels of spiked urine samples. The results are shown in Table 2; the accuracy, expressed as a percentage of recovery was within the range of 91.1%–106.9%, and the relative standard deviations were less than 6.4%. These results indicate that the method had good accuracy and precision, which was enough validity of the proposed method for the quantitative determination of sarcosine in real samples. The proposed method was compared with other reported methods34–38 for the detection of sarcosine, and the results are listed in Table S2.† This method showed better selectivity and similar sensitivity.
| Spiked (μg mL−1) | Found (μg mL−1) | Recovery (%) | RSD (%) |
|---|---|---|---|
| 0.5 | 0.53 ± 0.03 | 106.9 | 5.6 |
| 1 | 0.94 ± 0.06 | 94.1 | 6.4 |
| 5 | 4.55 ± 0.18 | 91.1 | 4.0 |
Experimental
Materials and regents
Horseradish peroxidase (HRP) was obtained from McLean Biochemical Reagent, Ltd (China). Glucose oxidase was purchased from Shanghai yuanye Bio-Technology Co., Ltd (China). Glucose and FeCl3·6H2O were purchased from Tianjin Komiou Chemical Reagent Co., Ltd. Ovalbumin (OVA) was purchased from Aladdin (China). Bovine hemoglobin (BHB) was purchased from the National Institutes for Food and Drug Control (China). Human serum albumin (HSA), bovine serum albumin (BSA) and gelatin were obtained from Sigma-Aldrich. 4-Formylphenylboronic acid (FPBA), sodium cyanoborohydride, cytochrome C (Cyt), sarcosine, sarcosine oxidase, tetraethyl orthosilicate (TEOS), 3-aminopropyltriethoxysilane (APTES), lysozyme (Lyz), bromelain (Bro) and 3,3′,5,5′-tetramethylbenzidine (TMB) were purchased from J&K Scientific, Ltd (China). Tween-20 was purchased from Beyotime Biotechnology, Ltd (China). Ethylene glycol was supplied by Tianjin Fuyu Fine Chemical (China). Ultrapure water was purified by Molelement 1810b (China). All other chemicals were analytical reagent grade and provided by local suppliers.Synthesis of boric acid-modified Fe3O4 nanoparticles
The boric acid-modified Fe3O4 nanoparticles were synthesized according to a reported method,27 which has been synthesized in two steps: the synthesis of the amino-functionalized Fe3O4 nanospheres and amino-functionalized Fe3O4 was immobilized with boronic acid moieties. Briefly, 1.0 g FeCl3·6H2O, 2.0 g anhydrous sodium acetate and 6.5 g 1,6-hexaneamine were successively dispersed in 30 mL glycol under vigorous stirring. The mixture was continuously stirred at room temperature until the mixture changed into a transparent solution. Then, the above solution was transferred into an autoclave and the autoclaves were placed in a heating box at 200 °C for 6 h. After that, the mixture was cooled to room temperature. The resultant products were collected by magnetic separation, and washed with ethanol and deionized water several times to remove unreacted reagents, and finally dried at 50 °C. Subsequently, the amino-functionalized Fe3O4 nanospheres (500 mg) and FPBA (20 mg) were dispersed into an anhydrous methanol solution (20 mL) with ultrasonication to get homogeneous dispersion. The obtained homogeneous solution was continuously shaken at room temperature for 18 h, and the resultant products were washed with anhydrous methanol three times. After that, the resultant products and sodium cyanoborohydride (20 mg) were dispersed in 20 mL anhydrous methanol again, and the mixtures were continuously shaken at room temperature for 22 h. Finally, the products were washed with methanol and deionized water, and dried at 50 °C.Preparation of MIPs and NIPs
The MIPs were prepared as follows: boric acid-modified Fe3O4 nanoparticles (10 mg) were dispersed in a phosphate buffer solution (1 mL, pH 8.5) containing HRP (500 μg), and the mixtures were shaken at room temperature for 1 h. Then, the products were obtained by magnet and washed with phosphate buffer solution three times to remove un-immobilized template. Subsequently, the phosphate buffer solution was replaced with a solvent of 0.05% tween-20 (5 mL). Afterwards, TEOS (10 μL) and APTES (2.5 μL) were added and shaken for 16 h at room temperature. Finally, the resultant products were washed with ethanol and deionized water to remove the unreacted monomer, and then washed with a mixed solution of acetonitrile (30%) and acetic acid (5%) to extract the template protein. The MIPs were blocked by BSA (2 mg mL−1) to decrease the nonspecific binding groups before use. As a control, the non-imprinted polymers (NIPs) were also prepared using the same procedure mentioned above in the absence of the template.Characterization
The magnetic properties were measured with an MPMS-SQUID VSM-094 vibrating sample magnetometer (VSM) (MPMS, USA). The surface elements of the magnetic nanoparticles were analyzed by X-ray photoelectron spectroscopy (XPS) (AXIS, UK). The crystal structure of the magnetic nanomaterials was characterized by X-ray diffraction (XRD) (Shimadzu, Japan). The morphologies of the magnetic nanoparticles were observed by field emission scanning electron microscopy (SEM) (Hitachi, Japan) and an H-7650 field emission transmission electron microscope (TEM) (Hitachi, Japan). FT-IR spectra were recorded on a Nicolet iS50 spectrometer (Thermo Scientific, USA). Thermogravimetric analysis (TGA) of the obtained nanoparticles was performed with a TGA/DSC 3+ thermogravimetric analyzer (METTLER TOLEDO, Switzerland).Selectivity
The selectivity was investigated using human serum albumin, ovalbumin, cytochrome C, lysozyme and bromelain as reference compounds. 20 μg of MIPs was added into 150 μL of PBST buffer solution containing HRP (10 μg mL−1) and the same concentration of reference proteins. The reference proteins and horseradish peroxidase bound with MIPs competitively for 30 min at room temperature. The MIPs were separated with an external magnetic field and washed three times with PBST solutions. Then, 150 μL of the TMB reagent was added and incubated for 30 min, and the coloration reaction was terminated by the addition of 50 μL of H2SO4 (2 mol L−1). Finally, the catalytic result of an absorbance was measured using an enzymatic reader (Thermo Scientific, USA). The selectivity of MIPs was assessed by comparing with the interference degree of the added reference proteins.Preparation and evaluation of immobilized enzyme
2 mg of MIPs were dispersed into 2 mL of PBST solution with 500 μg mL−1 of HRP. The solution was incubated at room temperature for 4 h. After washing the unbound substances, the immobilized enzyme was dispersed again and stored for further use. The Michaelis–Menten constant (Km) values were used to evaluate the activity of the free and immobilized enzymes, and they were obtained according to the Lineweaver–Burk equation. Enzymatic reactions were performed as the following steps: each reaction solution consisted of a 1.5 mL sodium citrate buffer solution containing H2O2 (0.018%), 10 μL of the free enzyme or the immobilized enzyme (20 μg) and various concentrations of TMB (1.5 mL). The mixture was incubated at room temperature for 10 min, and then the absorbance was scanned by UV spectrograph (PERSEE TU-1810, China).Immobilized horseradish peroxidase from horseradish
A practical application of the MIPs was demonstrated by immobilizing HRP from a crude extract of horseradish. The enzyme immobilization was performed by the following steps: fresh horseradish (1 g) was homogenized and extracted with PBST solution (10 mL) for 4 h at 4 °C. Then, the homogenate of horseradish was filtered through paper and diluted 50-fold before use. Then, 20 μg of MIPs or NIPs were added to the resulting solution; after incubating for 30 min, the enzyme-catalyzed reaction was operated using the same procedure mentioned above.Analysis of glucose in plasma
Glucose detection was realized as follows: at first, 100 μL TMB solution and sodium citrate buffer solution (100 μL) containing different concentrations of glucose were mixed. Then, 20 μg immobilized enzyme and a 10 μL volume of glucose oxidase solution (1 mg mL−1) were added into the resulting solution. After reacting for 60 min, 50 μL H2SO4 (2 mol L−1) was added to the above mixture to terminate the catalytic reaction. Finally, the catalytic result of an absorbance was measured using an enzymatic reader. The corresponding absorbance values at different concentrations were used for the quantification of glucose. The plasma samples, obtained from the First Affiliated Hospital of Xi'an Jiaotong University (Xi'an, China), were diluted 50-fold to fall within the calibration range before analysis and operated using the same procedure mentioned above. Three concentration levels (50 μg mL−1, 25 μg mL−1 and 0.5 μg mL−1) of spiked plasma samples were used to evaluate the method accuracy and precision. The selectivity analysis for glucose detection experiments used vitamin C, sucrose, glycine and mannitol as reference compounds, and their concentrations were 10 times more than that of glucose.Analysis of sarcosine in urine
Sarcosine detection was realized as follows: a 10 μL volume of a sarcosine oxidase solution (1 mg mL−1) was added to a PBS buffer solution (100 μL) containing different concentrations of sarcosine, and the mixtures were maintained at room temperature for 30 min. Then, 20 μg immobilized enzyme and 100 μL TMB solution were added into the resulting solution; after reacting for 30 min, 50 μL H2SO4 (2 mol L−1) was added to the above mixture to terminate the catalytic reaction. Finally, the catalytic result of an absorbance was measured using an enzymatic reader. The corresponding absorbance values at different concentrations were used for the quantification of sarcosine. The urine samples, obtained from the First Affiliated Hospital of Xi'an Jiaotong University (Xi'an, China), were diluted 20-fold before use and operated using the same procedure mentioned above. Three concentration levels (5 μg mL−1, 1 μg mL−1, and 0.5 μg mL−1) of spiked urine samples were used to evaluate the method accuracy and precision. The selectivity analysis for the sarcosine detection experiments were performed using glucose, vitamin C, glycine, phenylalanine and arginine as reference compounds, and their concentration was 10 times more than that of sarcosine.Conclusions
In this study, novel magnetic MIPs based on boronate affinity were successfully synthesized by a silane emulsion self-assembly method for the recognition of HRP. The boronate affinity MIPs not only exhibited great magnetic properties for rapid separation, but also showed excellent selectivity towards the template of HRP. The resultant boronate affinity MIPs can be used to immobilize horseradish peroxidase from a crude extract of horseradish, avoiding the complicated purification process of enzymes. The immobilized HRP can be employed to establish visual methods for detecting glucose and sarcosine. The developed methods showed excellent sensitivity, specificity, accuracy and reliability. This work demonstrated the practicality of fabricating boronate affinity MIPs by the silane emulsion self-assembly method for the recognition of glycoproteins, and these polymers would have a promising application prospect in the specific recognition and facile immobilization of enzymes in complex extract samples. While the method presented here was for a typical representative of HRP, the generic nature of this method suggests that more MIPs for the immobilization of other enzymes can be produced using this method.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (Grant No. 81773689, 81573391, 81603073 and 81603370), and the Natural Science Foundation of Shaanxi Province (No. 2017JQ8026).Notes and references
- A. Amine, H. Mohammadi, I. Bourais and G. Palleschi, Biosens. Bioelectron., 2006, 21, 1405–1423 CrossRef CAS.
- J. Adamski, P. Nowak and J. Kochana, Electrochim. Acta, 2010, 55, 2363–2367 CrossRef CAS.
- L. El Harrad, I. Bourais, H. Mohammadi and A. Amine, Sensors, 2018, 18, 1–25 CrossRef.
- N. Kharrat, Y. B. Ali, S. Marzouk, Y.-T. Gargouri and M. Karra-Châabouni, Process Biochem., 2011, 46, 1083–1089 CrossRef CAS.
- R. Das, M. Talat, O. N. Srivastava and A. M. Kayastha, Food Chem., 2018, 245, 488–499 CrossRef CAS.
- A. Elagli, K. Belhacene, C. Vivien, P. Dhulster, R. Froidevaux and P. Supiot, J. Mol. Catal. B: Enzym., 2014, 110, 77–86 CrossRef CAS.
- H. Liang, S. Sun, Y. Zhou and Y. Liu, Catalysts, 2017, 7, 1–11 CAS.
- G. N. Chen, P. Q. Guo, Y. Wang, L. Wang, H. Shu, Y. Z. Li, W. H. Jing, C. Chang and Q. Fu, Talanta, 2019, 198, 55–62 CrossRef CAS.
- X. Xu, R. Liu, P. Guo, Z. Luo, X. Cai, H. Shu, Y. Ge, C. Chang and Q. Fu, Food Chem., 2018, 256, 91–97 CrossRef CAS.
- X. Xu, P. Guo, Z. Luo, Y. Ge, Y. Zhou, R. Chang, W. Du, C. Chang and Q. Fu, RSC Adv., 2017, 7, 18765–18774 RSC.
- X. T. Ma, X. W. He, W. Y. Li and Y. K. Zhang, Sens. Actuators, B, 2017, 246, 879–886 CrossRef CAS.
- Z. Wang, T. Qiu, L. Guo, J. Ye, L. He and X. Li, Chem. Eng. J., 2018, 332, 409–418 CrossRef CAS.
- J. Erdőssy, V. Horváth, A. Yarman, F. W. Scheller and R. E. Gyurcsányi, Trends Analyt. Chem., 2016, 79, 179–190 CrossRef.
- X. Bi and Z. Liu, Anal. Chem., 2014, 86, 12382–12389 CrossRef CAS.
- X. Tu, P. Muhammad, J. Liu, Y. Ma, S. Wang, D. Yin and Z. Liu, Anal. Chem., 2016, 88, 12363–12370 CrossRef CAS.
- M. You, S. Yang, W. Tang, F. Zhang and P. G. He, ACS Appl. Mater. Interfaces, 2017, 9, 13855–13864 CrossRef CAS PubMed.
- P. Muhammad, X. Tu, J. Liu, Y. Wang and Z. Liu, ACS Appl. Mater. Interfaces, 2017, 9, 12082–12091 CrossRef CAS.
- J. Ye, Y. Chen and Z. Liu, Angew. Chem., Int. Ed., 2014, 53, 10386–10389 CrossRef CAS.
- Z. Wang, Y. Zhang, B. Zhang and X. Lu, Talanta, 2018, 190, 1–8 CrossRef CAS.
- L. Shang, C. Liu, M. Watanabe, B. Chen and K. Hayashi, Sens. Actuators, B, 2017, 249, 14–21 CrossRef CAS.
- J. Wang, M. Gu, J. Di, Y. Gao, Y. Wu and Y. Tu, Bioprocess Biosyst. Eng., 2007, 30, 289–296 CrossRef CAS.
- M. Zhao, Y. Li, X. Ma, M. Xia and Y. Zhang, Talanta, 2019, 200, 293–299 CrossRef CAS.
- A. Sreekumar, L. M. Poisson, T. M. Rajendiran, A. P. Khan, Q. Cao, J. Yu, B. Laxman, R. Mehra, R. J. Lonigro, Y. Li, M. K. Nyati, A. Ahsan, S. Kalyana-Sundaram, B. Han, X. Cao, J. Byun, G. S. Omenn, D. Ghosh, S. Pennathur, D. C. Alexander, A. Berger, J. R. Shuster, J. T. Wei, S. Varambally, C. Beecher and A. M. Chinnaiyan, Nature, 2009, 457, 910–914 CrossRef CAS.
- C. Valeri, P. Pozzilli and D. Leslie, Diabetes Metab. Res. Rev., 2004, 20(Suppl 2), S1–S8 CrossRef CAS.
- J. Lan, W. Xu, Q. Wan, X. Zhang, J. Lin, J. Chen and J. Chen, Anal. Chim. Acta, 2014, 825, 63–68 CrossRef CAS.
- Y. Pan, Y. Pang, Y. Shi, W. Zheng, Y. Long, Y. Huang and H. Zheng, Microchim. Acta, 2019, 186, 1–8 CrossRef CAS.
- M. Peng, H. Xiang, X. Hu, S. Shi and X. Chen, J. Chromatogr. A, 2016, 1474, 8–13 CrossRef CAS.
- Q. Chen, J. Chen, C. Gao, M. Zhang, J. Chen and H. Qiu, Analyst, 2015, 140, 2857–2863 RSC.
- M. M. Liu, X. Lian, H. Liu, Z. Z. Guo, H. H. Huang, Y. Lei, H. P. Peng, W. Chen, X. H. Lin, A. L. Liu and X. H. Xia, Talanta, 2019, 200, 511–517 CrossRef CAS.
- N. Alizadeh, A. Salimi and R. Hallaj, Sens. Actuators, B, 2019, 288, 44–52 CrossRef CAS.
- M. Liu, R. Liu and W. Chen, Biosens. Bioelectron., 2013, 45, 206–212 CrossRef CAS.
- J. L. Ma, B. C. Yin, X. Wu and B. C. Ye, Anal. Chem., 2017, 89, 1323–1328 CrossRef CAS.
- T. Zhu, Y. Zhang, L. Luo and X. Zhao, ACS Appl. Mater. Interfaces, 2019, 11, 10856–10861 CrossRef CAS.
- Q. Wang, C. Yang, Q. Yang, S. Yu and H. Yang, Anal. Chim. Acta, 2019, 1046, 93–98 CrossRef CAS.
- Y. Luo, J. Wang, L. Yang, T. Gao and R. Pei, Sens. Actuators, B, 2018, 276, 128–135 CrossRef CAS.
- Z. Xue, B. Yin, H. Wang, M. Li, H. Rao, X. Liu, X. Zhou and X. Lu, Nanoscale, 2016, 8, 5488–5496 RSC.
- O. Zitka, Z. Heger, M. Kominkova, S. Skalickova, S. Krizkova, V. Adam and R. Kizek, J. Sep. Sci., 2014, 37, 465–575 CrossRef CAS.
- T. Liu, B. Fu, J. Chen and K. Li, Microchim. Acta, 2019, 186, 1–8 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c9an01777a |
| This journal is © The Royal Society of Chemistry 2020 |